Example intronic APA qtls and eQTLs
Briana Mittleman
5/21/2020
Last updated: 2020-05-21
Checks: 7 0
Knit directory: apaQTL/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.6.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20190411) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: code/.Rhistory
Ignored: data/.DS_Store
Ignored: data/ProSeq/
Ignored: output/.DS_Store
Untracked files:
Untracked: .Rprofile
Untracked: ._.DS_Store
Untracked: .gitignore
Untracked: @
Untracked: GEO_brimittleman/
Untracked: _workflowr.yml
Untracked: analysis/._PASdescriptiveplots.Rmd
Untracked: analysis/._cuttoffPercUsage.Rmd
Untracked: analysis/APApeak_Phenotype_GeneLocAnno.Nuclear.5perc.fc.gz.qqnorm.allChrom
Untracked: analysis/APApeak_Phenotype_GeneLocAnno.Total.5perc.fc.gz.qqnorm.allChrom
Untracked: analysis/QTLexampleplots.Rmd
Untracked: analysis/cuttoffPercUsage.Rmd
Untracked: analysis/eQTLoverlap.Rmd
Untracked: analysis/interpret verify bam.Rmd
Untracked: analysis/interpret_verifybam.Rmd
Untracked: analysis/mergeRNA.Rmd
Untracked: analysis/oldstuffNotNeeded.Rmd
Untracked: analysis/remove_badlines.Rmd
Untracked: analysis/totSpecInNuclear.Rmd
Untracked: analysis/totSpecIncludenotTested.Rmd
Untracked: analysis/totalspec.Rmd
Untracked: analysis/usageCorr.Rmd
Untracked: apaQTL.Rproj
Untracked: checksumsfastq.txt.gz
Untracked: code/.NascentRNAdtPlotFirstintronicPAS.sh.swp
Untracked: code/._Allsplicesite2fasta.py
Untracked: code/._ApaQTL_nominalNonnorm.sh
Untracked: code/._BothFracDTPlotGeneRegions.sh
Untracked: code/._BothFracDTPlotGeneRegions_normalized.sh
Untracked: code/._ClosestTissuePAS.sh
Untracked: code/._ColocApAeQTL.sh
Untracked: code/._ColocApAeQTL_PM.sh
Untracked: code/._Coloc_generalAPAeQTL.R
Untracked: code/._Coloc_generalAPAeQTL_PM.R
Untracked: code/._CreateRNALZforeQTLs.sh
Untracked: code/._CreateRNALZnucAPAqtls.sh
Untracked: code/._DistPAS2Sig_RandomIntron.py
Untracked: code/._EandPqtl_perm.sh
Untracked: code/._EandPqtls.sh
Untracked: code/._ExtractGene4eQTLLZ.py
Untracked: code/._ExtractGene4eQTLLZpy
Untracked: code/._ExtractGeneRNAAssoc.py
Untracked: code/._ExtractPAS4LZeQTLs.py
Untracked: code/._ExtractPAS4eQTLsLZ.sh
Untracked: code/._ExtractPASforLZ.py
Untracked: code/._ExtractPASforLZ_run.sh
Untracked: code/._FC_NucintornUpandDown.sh
Untracked: code/._FC_UTR.sh
Untracked: code/._FC_intornUpandDownsteamPAS.sh
Untracked: code/._FC_nascentseq.sh
Untracked: code/._FC_newPeaks_olddata.sh
Untracked: code/._HMMpermuteTotal.py
Untracked: code/._HmmPermute.py
Untracked: code/._IntronicPASDT.sh
Untracked: code/._LC_samplegroups.py
Untracked: code/._LD_qtl.sh
Untracked: code/._LD_snpsproxy.sh
Untracked: code/._MapAllRBP.sh
Untracked: code/._NascentRNAdtPlot.sh
Untracked: code/._NascentRNAdtPlot3UTRPAS.sh
Untracked: code/._NascentRNAdtPlotExcludeFirstintronicPAS.sh
Untracked: code/._NascentRNAdtPlotNucPAS.sh
Untracked: code/._NascentRNAdtPlotTotPAS.sh
Untracked: code/._NascentRNAdtPlotintronicPAS.sh
Untracked: code/._NascnetRNAdtPlotPAS.sh
Untracked: code/._NetSeq_fourthintronDT.sh
Untracked: code/._NomResfromPASSNP.py
Untracked: code/._NuclearPAS_5per.bed.py
Untracked: code/._NuclearandRNA5samp_dtplots.sh
Untracked: code/._PTTfacetboxplots.R
Untracked: code/._PrematureQTLNominal.sh
Untracked: code/._PrematureQTLPermuted.sh
Untracked: code/._QTL2bed.py
Untracked: code/._QTL2bed_withstrand.py
Untracked: code/._RBPdisrupt.sh
Untracked: code/._RNAbam2bw.sh
Untracked: code/._RNAseqDTplot.sh
Untracked: code/._Randomsplicesite2fasta.py
Untracked: code/._Rplots.pdf
Untracked: code/._RunRes2PAS.sh
Untracked: code/._SAF215upbed.py
Untracked: code/._SnakefilePAS
Untracked: code/._SnakefilefiltPAS
Untracked: code/._TESplots100bp.sh
Untracked: code/._TESplots150bp.sh
Untracked: code/._TESplots200bp.sh
Untracked: code/._TotalPAS_5perc.bed.py
Untracked: code/._Totspec_example.sh
Untracked: code/._Totspec_exampleTOT.sh
Untracked: code/._Untitled
Untracked: code/._ZipandTabPheno.sh
Untracked: code/._aAPAqtl_nominal39ind.sh
Untracked: code/._allNucSpecQTLine.py
Untracked: code/._allNucSpecfromNonNorm.py
Untracked: code/._annotatePacBioPASregion.sh
Untracked: code/._annotatedPAS2bed.py
Untracked: code/._apaInPandE.py
Untracked: code/._apaQTLCorrectPvalMakeQQ.R
Untracked: code/._apaQTLCorrectpval_6or7a.R
Untracked: code/._apaQTL_Nominal.sh
Untracked: code/._apaQTL_nominalInclusive.sh
Untracked: code/._apaQTL_nominalv67.sh
Untracked: code/._apaQTL_permuted.sh
Untracked: code/._apaQTL_permuted_test6A7A.sh
Untracked: code/._apainRibo.py
Untracked: code/._assignNucIntonpeak2intronlocs.sh
Untracked: code/._assignTotIntronpeak2intronlocs.sh
Untracked: code/._bam2BW_5primemost.sh
Untracked: code/._bed2saf.py
Untracked: code/._bothFracDTplot1stintron.sh
Untracked: code/._bothFracDTplot4thintron.sh
Untracked: code/._bothFrac_FC.sh
Untracked: code/._callPeaksYL.py
Untracked: code/._changeRibonomQTLres2genename.py
Untracked: code/._changenomQTLres2geneName.py
Untracked: code/._chooseAnno2PAS_pacbio.py
Untracked: code/._chooseAnno2SAF.py
Untracked: code/._chooseSignalSite
Untracked: code/._chooseSignalSite.py
Untracked: code/._closestannotated.sh
Untracked: code/._closestannotated_byfrac.sh
Untracked: code/._cluster.json
Untracked: code/._clusterPAS.json
Untracked: code/._clusterfiltPAS.json
Untracked: code/._codingdms2bed.py
Untracked: code/._config.yaml
Untracked: code/._config2.yaml
Untracked: code/._configOLD.yaml
Untracked: code/._convertNominal2SNPLOC.py
Untracked: code/._convertNominal2SNPloc2Versions.py
Untracked: code/._convertNumeric.py
Untracked: code/._correctNomeqtl.R
Untracked: code/._createPlinkSampfile.py
Untracked: code/._dag.pdf
Untracked: code/._eQTL_switch2snploc.py
Untracked: code/._eQTLgenestestedapa.py
Untracked: code/._encodeRNADTplots.sh
Untracked: code/._extactPAS100meanphyloP.py
Untracked: code/._extractGeneLZfiles.sh
Untracked: code/._extractGeneLZfileseQTLs.sh
Untracked: code/._extractGenotypes.py
Untracked: code/._extractPACmeanPhyloP.py
Untracked: code/._extractPhylop50up.py
Untracked: code/._extractPhylopextra50.py
Untracked: code/._extractRNApval4lz.py
Untracked: code/._extractseqfromqtlfastq.py
Untracked: code/._fc2leafphen.py
Untracked: code/._fc_filteredPAS6and7As.sh
Untracked: code/._fifteenBPupstreamPAS.py
Untracked: code/._fiftyBPupstreamPAS.py
Untracked: code/._filter5perc.R
Untracked: code/._filter5percPheno.py
Untracked: code/._filterLDsnps.py
Untracked: code/._filterMPPAS.py
Untracked: code/._filterMPPAS_15.py
Untracked: code/._filterMPPAS_15_7As.py
Untracked: code/._filterMPPAS_50.py
Untracked: code/._filterSAFforMP.py
Untracked: code/._filterpeaks.py
Untracked: code/._finalPASbed2SAF.py
Untracked: code/._fix4su304corr.py
Untracked: code/._fix4su604corr.py
Untracked: code/._fix4sukalisto.py
Untracked: code/._fixExandUnexeQTL
Untracked: code/._fixExandUnexeQTL.py
Untracked: code/._fixFChead.py
Untracked: code/._fixFChead_bothfrac.py
Untracked: code/._fixFChead_short.py
Untracked: code/._fixGWAS4Munge.py
Untracked: code/._fixH3k12ac.py
Untracked: code/._fixPASregionSNPs.py
Untracked: code/._fixRNAhead4corr.py
Untracked: code/._fixRNAkalisto.py
Untracked: code/._fix_randomIntron.py
Untracked: code/._fixgroupedtranscript.py
Untracked: code/._fixhead_netseqfc.py
Untracked: code/._getAPAfromanyeQTL.py
Untracked: code/._getApapval4eqtl.py
Untracked: code/._getApapval4eqtl_unexp.py
Untracked: code/._getApapval4eqtl_version67.py
Untracked: code/._getDownstreamIntronNuclear.py
Untracked: code/._getIntronDownstreamPAS.py
Untracked: code/._getIntronUpstreamPAS.py
Untracked: code/._getQTLalleles.py
Untracked: code/._getQTLfastq.sh
Untracked: code/._getUpstreamIntronNuclear.py
Untracked: code/._grouptranscripts.py
Untracked: code/._intersectVCFandupPAS.sh
Untracked: code/._keep5perMAF.py
Untracked: code/._keepSNP_vcf.sh
Untracked: code/._make5percPeakbed.py
Untracked: code/._makeFileID.py
Untracked: code/._makePheno.py
Untracked: code/._makeSAFbothfrac5perc.py
Untracked: code/._makeSNP2rsidfile.py
Untracked: code/._makeeQTLempirical_unexp.py
Untracked: code/._makeeQTLempiricaldist.py
Untracked: code/._makegencondeTSSfile.py
Untracked: code/._mapSSsnps2PAS.sh
Untracked: code/._mergRNABam.sh
Untracked: code/._mergeAllBam.sh
Untracked: code/._mergeAnnotations.sh
Untracked: code/._mergeBW_norm.sh
Untracked: code/._mergeBamNascent.sh
Untracked: code/._mergeByFracBam.sh
Untracked: code/._mergePeaks.sh
Untracked: code/._miRNAdisrupt.sh
Untracked: code/._mnase1stintron.sh
Untracked: code/._mnaseDT_fourthintron.sh
Untracked: code/._namePeaks.py
Untracked: code/._netseqDTplot1stIntron.sh
Untracked: code/._netseqFC.sh
Untracked: code/._nominavalfortotspec.py
Untracked: code/._noninalpval4alltot.py
Untracked: code/._nucQTLGWAS.py
Untracked: code/._nucSpecQTLineData.py
Untracked: code/._nucSpeceffectsize.py
Untracked: code/._nucspecnucPASine.py
Untracked: code/._pQTLsotherdata.py
Untracked: code/._pacbioDT.sh
Untracked: code/._pacbioIntronicDT.sh
Untracked: code/._parseALLSSres.py
Untracked: code/._parseBestbamid.py
Untracked: code/._parseLDRes.py
Untracked: code/._parseLDresBothPAS.sh
Untracked: code/._parseRanodmSSres.py
Untracked: code/._parseSSres.py
Untracked: code/._peak2PAS.py
Untracked: code/._peakFC.sh
Untracked: code/._pheno2countonly.R
Untracked: code/._phenoQTLfromlist.py
Untracked: code/._processYRIgen.py
Untracked: code/._pttQTLsinapaQTL.py
Untracked: code/._qtlRegionseq.sh
Untracked: code/._qtlsPvalOppFrac.py
Untracked: code/._quantassign2parsedpeak.py
Untracked: code/._removeXfromHmm.py
Untracked: code/._removeloc_pheno.py
Untracked: code/._riboQTL.sh
Untracked: code/._runCorrectNomEqtl.sh
Untracked: code/._runFixGWAS4Munge.sh
Untracked: code/._runHMMpermuteAPAqtls.sh
Untracked: code/._runHMMpermuteeQTLS.sh
Untracked: code/._runMakeEmpiricaleQTL_unexp.sh
Untracked: code/._runMakeeQTLempirical.sh
Untracked: code/._run_bam2bw_all3prime.sh
Untracked: code/._run_bam2bw_extra3.sh
Untracked: code/._run_bestbamid.sj
Untracked: code/._run_dist2sig_randomintron.sh
Untracked: code/._run_filtersnpLD.sh
Untracked: code/._run_getAPAfromeQTL_version6.7.sh
Untracked: code/._run_getApaPval4eqtl.sh
Untracked: code/._run_getapafromeQTL.py
Untracked: code/._run_getapafromeQTL.sh
Untracked: code/._run_getapapval4eqtl_unexp.sh
Untracked: code/._run_leafcutterDiffIso.sh
Untracked: code/._run_prxySNP.sh
Untracked: code/._run_pttfacetboxplot.sh
Untracked: code/._run_sepUsagephen.sh
Untracked: code/._run_sepgenobychrom.sh
Untracked: code/._run_verifybam.sh
Untracked: code/._selectNominalPvalues.py
Untracked: code/._sepUsagePhen.py
Untracked: code/._sepgenobychrom.py
Untracked: code/._snakemakePAS.batch
Untracked: code/._snakemakefiltPAS.batch
Untracked: code/._sortindexRNAbam.sh
Untracked: code/._specAPAinE.py
Untracked: code/._splicesite2fasta.py
Untracked: code/._submit-snakemakePAS.sh
Untracked: code/._submit-snakemakefiltPAS.sh
Untracked: code/._subsetAPAnotEorPgene.py
Untracked: code/._subsetAPAnotEorPgene_2versions.py
Untracked: code/._subsetAPAnotEorR.py
Untracked: code/._subsetApanoteGene.py
Untracked: code/._subsetApanoteGene_2versions.py
Untracked: code/._subsetNootherQTL.py
Untracked: code/._subsetUnexplainedeQTLs.py
Untracked: code/._subsetVCF_SS.sh
Untracked: code/._subsetVCF_noSSregions.sh
Untracked: code/._subsetVCF_upstreamPAS.sh
Untracked: code/._subset_diffisopheno.py
Untracked: code/._subsetpermAPAwithGenelist.py
Untracked: code/._subsetpermAPAwithGenelist_2versions.py
Untracked: code/._subsetvcf_otherreg.sh
Untracked: code/._subsetvcf_permSS.sh
Untracked: code/._subtrachfiveprimeUTR.sh
Untracked: code/._subtractExons.sh
Untracked: code/._subtractfiveprimeUTR.sh
Untracked: code/._tabixSNPS.sh
Untracked: code/._tenBPupstreamPAS.py
Untracked: code/._test.pdf
Untracked: code/._testVerifyBam.sh
Untracked: code/._tissuePAS2hg19.sh
Untracked: code/._totSeceffectsize.py
Untracked: code/._totspecinE.py
Untracked: code/._totspecqtlFacetBoxplots.sh
Untracked: code/._totspecqtlFacetBoxplotsTOT.sh
Untracked: code/._twentyBPupstreamPAS.py
Untracked: code/._utrdms2saf.py
Untracked: code/._vcf2bed.py
Untracked: code/._verifyBam18517N.sh
Untracked: code/._verifyBam18517T.sh
Untracked: code/._verifyBam19128N.sh
Untracked: code/._verifyBam19128T.sh
Untracked: code/._wrap_verifybam.sh
Untracked: code/._writePTTexamplecode.py
Untracked: code/._writePTTexamplecode.sh
Untracked: code/.pversion
Untracked: code/.snakemake/
Untracked: code/1
Untracked: code/APAqtl_nominal.err
Untracked: code/APAqtl_nominal.out
Untracked: code/APAqtl_nominal_39.err
Untracked: code/APAqtl_nominal_39.out
Untracked: code/APAqtl_nominal_inclusive.err
Untracked: code/APAqtl_nominal_inclusive.out
Untracked: code/APAqtl_nominal_nonNorm.err
Untracked: code/APAqtl_nominal_nonNorm.out
Untracked: code/APAqtl_nominal_versions67.err
Untracked: code/APAqtl_nominal_versions67.out
Untracked: code/APAqtl_permuted.err
Untracked: code/APAqtl_permuted.out
Untracked: code/APAqtl_permuted_versions67.err
Untracked: code/APAqtl_permuted_versions67.out
Untracked: code/Allsplicesite2fasta.py
Untracked: code/BothFracDTPlot1stintron.err
Untracked: code/BothFracDTPlot1stintron.out
Untracked: code/BothFracDTPlot4stintron.err
Untracked: code/BothFracDTPlot4stintron.out
Untracked: code/BothFracDTPlotGeneRegions.err
Untracked: code/BothFracDTPlotGeneRegions.out
Untracked: code/BothFracDTPlotGeneRegions_norm.err
Untracked: code/BothFracDTPlotGeneRegions_norm.out
Untracked: code/ClosestTissuePAS.sh
Untracked: code/ColocApAeQTL.err
Untracked: code/ColocApAeQTL.out
Untracked: code/ColocApAeQTL.sh
Untracked: code/ColocApAeQTLPM.err
Untracked: code/ColocApAeQTLPM.out
Untracked: code/ColocApAeQTL_PM.sh
Untracked: code/Coloc_generalAPAeQTL.R
Untracked: code/Coloc_generalAPAeQTL_PM.R
Untracked: code/CreateRNALZforeQTLs.sh
Untracked: code/CreateRNALZnucAPAqtls.sh
Untracked: code/DistPAS2Sig_RandomIntron.py
Untracked: code/EandPqtl.err
Untracked: code/EandPqtl.out
Untracked: code/EncodeRNADTPlotGeneRegions.err
Untracked: code/EncodeRNADTPlotGeneRegions.out
Untracked: code/ExtractGene4eQTLLZ.py
Untracked: code/ExtractGene4eQTLLZpy
Untracked: code/ExtractGeneRNAAssoc.py
Untracked: code/ExtractPAS4LZeQTLs.py
Untracked: code/ExtractPAS4eQTLsLZ.sh
Untracked: code/ExtractPASforLZ.py
Untracked: code/ExtractPASforLZ_run.sh
Untracked: code/FC_NucintronPASupandDown.err
Untracked: code/FC_NucintronPASupandDown.out
Untracked: code/FC_UTR.err
Untracked: code/FC_UTR.out
Untracked: code/FC_intronPASupandDown.err
Untracked: code/FC_intronPASupandDown.out
Untracked: code/FC_nascent.err
Untracked: code/FC_nascentout
Untracked: code/FC_newPAS_olddata.err
Untracked: code/FC_newPAS_olddata.out
Untracked: code/HmmPermute.p
Untracked: code/IntronicPASDT.err
Untracked: code/IntronicPASDT.out
Untracked: code/LD_vcftools.hap.out
Untracked: code/MapAllRBP.sh
Untracked: code/MapRBP.err
Untracked: code/MapRBP.out
Untracked: code/NascentDTPlotGeneRegions.err
Untracked: code/NascentDTPlotGeneRegions.out
Untracked: code/NascentDTPlotPAS.err
Untracked: code/NascentDTPlotPAS.out
Untracked: code/NascentDTPlotPAS_3utr.err
Untracked: code/NascentDTPlotPAS_3utr.out
Untracked: code/NascentDTPlotPAS_firstintron.err
Untracked: code/NascentDTPlotPAS_firstintron.out
Untracked: code/NascentDTPlotPAS_intron.err
Untracked: code/NascentDTPlotPAS_intron.out
Untracked: code/NascentDTPlotPAS_nuc.err
Untracked: code/NascentDTPlotPAS_nuc.out
Untracked: code/NascentDTPlotPAS_tot.err
Untracked: code/NascentDTPlotPAS_tot.out
Untracked: code/Nuclear_example.err
Untracked: code/Nuclear_example.out
Untracked: code/NuclearandRNA5samp_dtplots.sh
Untracked: code/NuclearandRNAFracDTPlotGeneRegions.err
Untracked: code/NuclearandRNAFracDTPlotGeneRegions.out
Untracked: code/PACbioDT.err
Untracked: code/PACbioDT.out
Untracked: code/PACbioDTitronic.err
Untracked: code/PACbioDTitronic.out
Untracked: code/Prematureqtl_nominal.err
Untracked: code/Prematureqtl_nominal.out
Untracked: code/Prematureqtl_permuted.err
Untracked: code/Prematureqtl_permuted.out
Untracked: code/RBPdisrupt.err
Untracked: code/RBPdisrupt.out
Untracked: code/RBPdisrupt.sh
Untracked: code/README.md
Untracked: code/RNABam2BW.err
Untracked: code/RNABam2BW.out
Untracked: code/RNAseqDTPlotGeneRegions.err
Untracked: code/RNAseqDTPlotGeneRegions.out
Untracked: code/Randomsplicesite2fasta.py
Untracked: code/Rplots.pdf
Untracked: code/TESplots100bp.err
Untracked: code/TESplots100bp.out
Untracked: code/TESplots150bp.err
Untracked: code/TESplots150bp.out
Untracked: code/TESplots200bp.err
Untracked: code/TESplots200bp.out
Untracked: code/Tissueclosestannotated.err
Untracked: code/Tissueclosestannotated.out
Untracked: code/Total_example.err
Untracked: code/Total_example.out
Untracked: code/Totspec_example.err
Untracked: code/Totspec_example.out
Untracked: code/Totspec_example.sh
Untracked: code/Totspec_exampleTOT.err
Untracked: code/Totspec_exampleTOT.out
Untracked: code/Totspec_exampleTOT.sh
Untracked: code/Untitled
Untracked: code/YRI_LCL.vcf.gz
Untracked: code/YRI_LCL_chr1.vcf.gz.log
Untracked: code/YRI_LCL_chr1.vcf.gz.recode.vcf
Untracked: code/annotatedPASregion.err
Untracked: code/annotatedPASregion.out
Untracked: code/apaQTL_nominalInclusive.sh
Untracked: code/assignPeak2Intronicregion.err
Untracked: code/assignPeak2Intronicregion.out
Untracked: code/assigntotPeak2Intronicregion.err
Untracked: code/assigntotPeak2Intronicregion.out
Untracked: code/bam2bw.err
Untracked: code/bam2bw.out
Untracked: code/bam2bw_5primemost.err
Untracked: code/bam2bw_5primemost.out
Untracked: code/binary_fileset.log
Untracked: code/bothFrac_FC.err
Untracked: code/bothFrac_FC.out
Untracked: code/callSHscripts.txt
Untracked: code/closestannotated.err
Untracked: code/closestannotated.out
Untracked: code/closestannotatedbyfrac.err
Untracked: code/closestannotatedbyfrac.out
Untracked: code/dag.pdf
Untracked: code/dagPAS.pdf
Untracked: code/dagfiltPAS.pdf
Untracked: code/extactPAS100meanphyloP.py
Untracked: code/extractGeneLZfiles.err
Untracked: code/extractGeneLZfiles.out
Untracked: code/extractGeneLZfiles.sh
Untracked: code/extractGeneLZfileseQTLs.err
Untracked: code/extractGeneLZfileseQTLs.out
Untracked: code/extractGeneLZfileseQTLs.sh
Untracked: code/extractPACmeanPhyloP.py
Untracked: code/extractPASLZfiles.err
Untracked: code/extractPASLZfiles.out
Untracked: code/extractPASLZfileseQTLs.err
Untracked: code/extractPASLZfileseQTLs.out
Untracked: code/extractPhylop50up.py
Untracked: code/extractPhylopextra50.py
Untracked: code/extractRNApval4lz.py
Untracked: code/fixExandUnexeQTL
Untracked: code/fixGWAS4Munge.py
Untracked: code/fix_randomIntron.py
Untracked: code/fixmunge
Untracked: code/genotypesYRI.gen.proc.keep.vcf.log
Untracked: code/genotypesYRI.gen.proc.keep.vcf.recode.vcf
Untracked: code/getseq100up.err
Untracked: code/getseq100up.out
Untracked: code/grouptranscripts.err
Untracked: code/grouptranscripts.out
Untracked: code/intersectPAS_ssSNPS.err
Untracked: code/intersectPAS_ssSNPS.out
Untracked: code/intersectVCFPAS.err
Untracked: code/intersectVCFPAS.out
Untracked: code/liftoverPAShg38to19.err
Untracked: code/liftoverPAShg38to19.out
Untracked: code/log/
Untracked: code/logs/
Untracked: code/merge53PRIMEbam.err
Untracked: code/merge53PRIMEbam.out
Untracked: code/merge53RNAbam.err
Untracked: code/merge53prime.sh
Untracked: code/merge5RNABam.err
Untracked: code/merge5RNABam.out
Untracked: code/merge5RNAbam.out
Untracked: code/merge5RNAbam.sh
Untracked: code/mergeAnno.err
Untracked: code/mergeAnno.out
Untracked: code/mergeBWnorm.err
Untracked: code/mergeBWnorm.out
Untracked: code/mergeBamNacent.err
Untracked: code/mergeBamNacent.out
Untracked: code/mergeRNAbam.err
Untracked: code/mergeRNAbam.out
Untracked: code/miRNAdisrupt.err
Untracked: code/miRNAdisrupt.out
Untracked: code/miRNAdisrupt.sh
Untracked: code/mnaseDTPlot1stintron.err
Untracked: code/mnaseDTPlot1stintron.out
Untracked: code/mnaseDTPlot4thintron.err
Untracked: code/mnaseDTPlot4thintron.out
Untracked: code/netDTPlot4thintron.out
Untracked: code/netseqFC.err
Untracked: code/netseqFC.out
Untracked: code/neyDTPlot4thintron.err
Untracked: code/nominavalfortotspec.py
Untracked: code/noninalpval4alltot.py
Untracked: code/nucspecinE.py
Untracked: code/parseALLSSres.py
Untracked: code/parseLDRes.py
Untracked: code/parseLDres.err
Untracked: code/parseLDres.out
Untracked: code/parseLDresBothPAS.sh
Untracked: code/parseRanodmSSres.py
Untracked: code/parseSSres.py
Untracked: code/plink.log
Untracked: code/prxySNP.err
Untracked: code/prxySNP.out
Untracked: code/pttFacetBoxplots.err
Untracked: code/pttFacetBoxplots.out
Untracked: code/qtlFacetBoxplots.err
Untracked: code/qtlFacetBoxplots.out
Untracked: code/rLD_vcftools.hap.err
Untracked: code/riboqtl.err
Untracked: code/riboqtl.out
Untracked: code/runBestBamID.err
Untracked: code/runCorrectNomeqtl.err
Untracked: code/runCorrectNomeqtl.out
Untracked: code/runFilterLD.err
Untracked: code/runFilterLD.out
Untracked: code/runFixGWAS4Munge.sh
Untracked: code/runHMMpermute.err
Untracked: code/runHMMpermute.out
Untracked: code/runHMMpermuteeQTLs.err
Untracked: code/runHMMpermuteeQTLs.out
Untracked: code/runMakeEmpiricaleQTLs.err
Untracked: code/runMakeEmpiricaleQTLs.out
Untracked: code/runMakeEmpiricaleQTLsunex.err
Untracked: code/runMakeEmpiricaleQTLsunex.out
Untracked: code/run_DistPAS2Sig.err
Untracked: code/run_DistPAS2Sig.out
Untracked: code/run_DistPAS2Sig_intron.err
Untracked: code/run_DistPAS2Sig_intron.out
Untracked: code/run_bam2bw.err
Untracked: code/run_bam2bw.out
Untracked: code/run_bam2bwexta.err
Untracked: code/run_bam2bwexta.out
Untracked: code/run_dist2sig_randomintron.sh
Untracked: code/run_getAPAfromanyeQTL.err
Untracked: code/run_getAPAfromanyeQTL.out
Untracked: code/run_getApaPval4eQTLs.err
Untracked: code/run_getApaPval4eQTLs.out
Untracked: code/run_getApaPval4eQTLsunexplained.err
Untracked: code/run_getApaPval4eQTLsunexplained.out
Untracked: code/run_leafcutter_ds.err
Untracked: code/run_leafcutter_ds.out
Untracked: code/run_sepgenobychrom.err
Untracked: code/run_sepgenobychrom.out
Untracked: code/run_sepusage.err
Untracked: code/run_sepusage.out
Untracked: code/run_verifybam.err
Untracked: code/run_verifybam.out
Untracked: code/run_verifybam128N.err
Untracked: code/run_verifybam128N.out
Untracked: code/run_verifybam128T.err
Untracked: code/run_verifybam128T.out
Untracked: code/run_verifybam517N.err
Untracked: code/run_verifybam517N.out
Untracked: code/run_verifybam517T.err
Untracked: code/run_verifybam517T.out
Untracked: code/runprxySNP.err
Untracked: code/runprxySNP.out
Untracked: code/runres2pas.err
Untracked: code/runres2pas.out
Untracked: code/scripts/
Untracked: code/scripts_PAS_500_Lymph/
Untracked: code/seqQTLfastq.err
Untracked: code/seqQTLfastq.out
Untracked: code/seqQTLregion.err
Untracked: code/seqQTLregion.out
Untracked: code/snakePASlog.out
Untracked: code/snakefiltPASlog.out
Untracked: code/sortindexRNABam.err
Untracked: code/sortindexRNABam.out
Untracked: code/specAPAinE.py
Untracked: code/splicesite2fasta.py
Untracked: code/subsetAPAnotEorR.py
Untracked: code/subsetNootherQTL.py
Untracked: code/subsetvcf_SS.err
Untracked: code/subsetvcf_SS.out
Untracked: code/subsetvcf_noSS.err
Untracked: code/subsetvcf_noSS.out
Untracked: code/subsetvcf_pas.err
Untracked: code/subsetvcf_pas.out
Untracked: code/subsetvcf_perm.err
Untracked: code/subsetvcf_perm.out
Untracked: code/subsetvcf_rand.err
Untracked: code/subsetvcf_rand.out
Untracked: code/subtract5UTR.err
Untracked: code/subtract5UTR.out
Untracked: code/subtractExons.err
Untracked: code/subtractExons.out
Untracked: code/tabixSNPs.err
Untracked: code/tabixSNPs.out
Untracked: code/test.pdf
Untracked: code/testFix.txt
Untracked: code/test_verifybam.err
Untracked: code/test_verifybam.out
Untracked: code/tissuePAS2hg19.sh
Untracked: code/totspecinE.py
Untracked: code/totspecqtlFacetBoxplots.err
Untracked: code/totspecqtlFacetBoxplots.out
Untracked: code/totspecqtlFacetBoxplots.sh
Untracked: code/totspecqtlFacetBoxplotsTOT.err
Untracked: code/totspecqtlFacetBoxplotsTOT.out
Untracked: code/totspecqtlFacetBoxplotsTOT.sh
Untracked: code/vcf_keepsnps.err
Untracked: code/vcf_keepsnps.out
Untracked: code/wrap_verifybam.err
Untracked: code/wrap_verifybam.out
Untracked: code/zipandtabPhen.err
Untracked: code/zipandtabPhen.out
Untracked: data/._.DS_Store
Untracked: data/._MetaDataSequencing.txt
Untracked: data/AnnotatedPAS/
Untracked: data/ApaByEgene/
Untracked: data/ApaByPgene/
Untracked: data/ApaByRgene/
Untracked: data/BadLines/
Untracked: data/BaseComp/
Untracked: data/Battle_pQTL/
Untracked: data/CheckSums/
Untracked: data/CompareOldandNew/
Untracked: data/DTmatrix/
Untracked: data/DiffIso/
Untracked: data/EncodeRNA/
Untracked: data/ExampleQTLPlots/
Untracked: data/ExampleQTLPlots_update/
Untracked: data/ExpressionIndependentapaQTLs.txt
Untracked: data/FiveMergedBW/
Untracked: data/FiveMergedBam/
Untracked: data/FlaggedPAS/
Untracked: data/GWAS_overlap/
Untracked: data/Geuvadis/
Untracked: data/GeuvadisRNA/
Untracked: data/GeuvadiseQTL/
Untracked: data/HMMqtls/
Untracked: data/LDSR_annotations/
Untracked: data/LZ_both/
Untracked: data/Li_eQTLs/
Untracked: data/NMD/
Untracked: data/NascentRNA/
Untracked: data/NucSpeceQTLeffect/
Untracked: data/PAS/
Untracked: data/PAS_postFlag/
Untracked: data/PolyA_DB/
Untracked: data/PreTerm_pheno/
Untracked: data/PrematureQTLNominal/
Untracked: data/PrematureQTLPermuted/
Untracked: data/QTLGenotypes/
Untracked: data/QTLoverlap/
Untracked: data/QTLoverlap_inclusive/
Untracked: data/QTLoverlap_nonNorm/
Untracked: data/README.md
Untracked: data/RNAseq/
Untracked: data/Reads2UTR/
Untracked: data/SNPinSS/
Untracked: data/SignalSiteFiles/
Untracked: data/TF_motifdisruption/
Untracked: data/TSS/
Untracked: data/ThirtyNineIndQtl_nominal/
Untracked: data/TissueData/
Untracked: data/Version15bp6As/
Untracked: data/Version15bp7As/
Untracked: data/apaQTLNominal/
Untracked: data/apaQTLNominal_4pc/
Untracked: data/apaQTLNominal_inclusive/
Untracked: data/apaQTLPermuted/
Untracked: data/apaQTLPermuted_4pc/
Untracked: data/apaQTLs/
Untracked: data/assignedPeaks/
Untracked: data/assignedPeaks_15Up/
Untracked: data/bam/
Untracked: data/bam_clean/
Untracked: data/bam_waspfilt/
Untracked: data/bed_10up/
Untracked: data/bed_clean/
Untracked: data/bed_clean_sort/
Untracked: data/bed_waspfilter/
Untracked: data/bedsort_waspfilter/
Untracked: data/bothFrac_FC/
Untracked: data/bw/
Untracked: data/bw_norm/
Untracked: data/coloc/
Untracked: data/coloc_PM/
Untracked: data/eCLip/
Untracked: data/eQTL_LZ/
Untracked: data/eQTLs/
Untracked: data/exampleQTLs/
Untracked: data/exosome/
Untracked: data/fastq/
Untracked: data/filterPeaks/
Untracked: data/fourSU/
Untracked: data/h3k27ac/
Untracked: data/highdiffsiggenes.txt
Untracked: data/inclusivePeaks/
Untracked: data/inclusivePeaks_FC/
Untracked: data/intronRNAratio/
Untracked: data/intron_analysis/
Untracked: data/locusZoom/
Untracked: data/mergedBG/
Untracked: data/mergedBW_byfrac/
Untracked: data/mergedBW_norm/
Untracked: data/mergedBam/
Untracked: data/mergedbyFracBam/
Untracked: data/miRNAbinding/
Untracked: data/molPhenos/
Untracked: data/molQTLs/
Untracked: data/motifdistrupt/
Untracked: data/nPAS/
Untracked: data/netseq/
Untracked: data/nonNorm_pheno/
Untracked: data/nuc_10up/
Untracked: data/nuc_10upclean/
Untracked: data/oldPASfiles/
Untracked: data/overlapeQTL_try2/
Untracked: data/overlapeQTLs/
Untracked: data/pQTLoverlap/
Untracked: data/pacbio/
Untracked: data/peakCoverage/
Untracked: data/peaks_5perc/
Untracked: data/phenotype/
Untracked: data/phenotype_5perc/
Untracked: data/phenotype_inclusivePAS/
Untracked: data/phylop/
Untracked: data/pttQTL/
Untracked: data/pttQTLplots/
Untracked: data/sigDiffGenes.txt
Untracked: data/sort/
Untracked: data/sort_clean/
Untracked: data/sort_waspfilter/
Untracked: data/splicesite/
Untracked: data/totSpecExampleQTLPlots/
Untracked: data/totSpecExampleQTLPlots_tot/
Untracked: data/twoMech/
Untracked: data/vareQTLvarAPAqtl/
Untracked: data/verifyBAM/
Untracked: data/verifyBAM_full/
Untracked: nohup.out
Untracked: output/._.DS_Store
Untracked: output/._AverageDiffHeatmap.Nuclear.png
Untracked: output/._AverageDiffHeatmap.Total.png
Untracked: output/._GeneswithAPApotential.png
Untracked: output/._GeneswithAPApotentialAllPAS.png
Untracked: output/._PASlocation.png
Untracked: output/._SignalSitePlot.png
Untracked: output/._meanCorrelationPhenotypes.svg
Untracked: output/._qqplot_Nuclear_APAperm.png
Untracked: output/._qqplot_Nuclear_APAperm_4pc.png
Untracked: output/._qqplot_Total_APAperm.png
Untracked: output/._qqplot_Total_APAperm_4pc.png
Untracked: output/AverageDiffHeatmap.Nuclear.png
Untracked: output/AverageDiffHeatmap.Total.png
Untracked: output/GeneswithAPApotential.png
Untracked: output/GeneswithAPApotentialAllPAS.png
Untracked: output/PASlocation.png
Untracked: output/SignalSitePlot.png
Untracked: output/SignalSitePlotbyLoc.png
Untracked: output/dtPlots/
Untracked: output/fastqc/
Untracked: output/meanCorrelationPhenotypes.svg
Untracked: output/newnuc.png
Untracked: output/newtot.png
Untracked: output/oldnuc.png
Untracked: output/oldtot.png
Untracked: output/qqplot_Nuclear_APAperm.png
Untracked: output/qqplot_Nuclear_APAperm_4pc.png
Untracked: output/qqplot_Total_APAperm.png
Untracked: output/qqplot_Total_APAperm_4pc.png
Untracked: run_verifybam517N.err
Untracked: run_verifybam517N.out
Unstaged changes:
Modified: analysis/NuclearSpecIncludeNotTested.Rmd
Modified: analysis/PASdescriptiveplots.Rmd
Modified: analysis/Readdistagainstfeatures.Rmd
Modified: analysis/TSS.Rmd
Modified: analysis/apabyeQTLstatus.Rmd
Modified: analysis/decayAndStability.Rmd
Modified: analysis/miRNAdisrupt.Rmd
Modified: analysis/nascenttranscription.Rmd
Modified: analysis/nucSpecinEQTLs.Rmd
Modified: analysis/overlapapaqtlsandeqtls.Rmd
Modified: analysis/pQTLexampleplot.Rmd
Modified: analysis/version15bpfilter.Rmd
Modified: code/DistPAS2Sig.py
Modified: code/Script4NuclearQTLexamples.sh
Modified: code/Script4TotalQTLexamples.sh
Modified: code/apaQTLsnake.err
Modified: code/apaqtlfacetboxplots.R
Modified: code/environment.yaml
Modified: code/run_qtlFacetBoxplots.sh
Deleted: code/test.txt
Deleted: reads_graphs.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | a67b63a | brimittleman | 2020-05-21 | verify dir |
library(tidyverse)── Attaching packages ──────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.2.1 ✔ purrr 0.3.2
✔ tibble 2.1.3 ✔ dplyr 0.8.3
✔ tidyr 0.8.3 ✔ stringr 1.4.0
✔ readr 1.3.1 ✔ forcats 0.4.0── Conflicts ─────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()Check to make sure effect size are the same direction. Check top examples from the eQTL vs intronic apaQTL plot.
eQTLeffect=read.table("../data/molQTLs/fastqtl_qqnorm_RNAseq_phase2.fixed.nominal.AllNomRes.GeneName_snploc.txt", stringsAsFactors = F, col.names = c("gene","snp","dist", "pval", "eQTL_es")) %>% select(gene, snp, eQTL_es)nomnames=c("peakID", 'snp','dist', 'pval', 'slope')
nuclearapaUnexplained=read.table("../data/overlapeQTL_try2/apaNuclear_unexplainedQTLs.txt", stringsAsFactors = F, col.names = nomnames) %>% separate(peakID, into=c("chr","start","end","geneID"), sep=":") %>% separate(geneID, into=c("gene", "loc", "strand", "PASnum"), sep="_") %>% group_by(gene, snp) %>% mutate(nPeaks=n(), adjPval=pval* nPeaks) %>% dplyr::slice(which.min(adjPval))
nuclearapaexplained=read.table("../data/overlapeQTL_try2/apaNuclear_explainedQTLs.txt", stringsAsFactors = F, col.names = nomnames) %>% separate(peakID, into=c("chr","start","end","geneID"), sep=":") %>% separate(geneID, into=c("gene", "loc", "strand", "PASnum"), sep="_") %>% group_by(gene, snp) %>% mutate(nPeaks=n(), adjPval=pval* nPeaks) %>% dplyr::slice(which.min(adjPval))
alleQTLS_nuclear=bind_rows(nuclearapaUnexplained,nuclearapaexplained) %>% filter(loc=="intron") %>% inner_join(eQTLeffect, by=c("gene","snp"))ggplot(alleQTLS_nuclear,aes(x=eQTL_es, y=slope)) + geom_point() + geom_smooth(method = "lm") + geom_text(aes(label=gene), nudge_y = .1)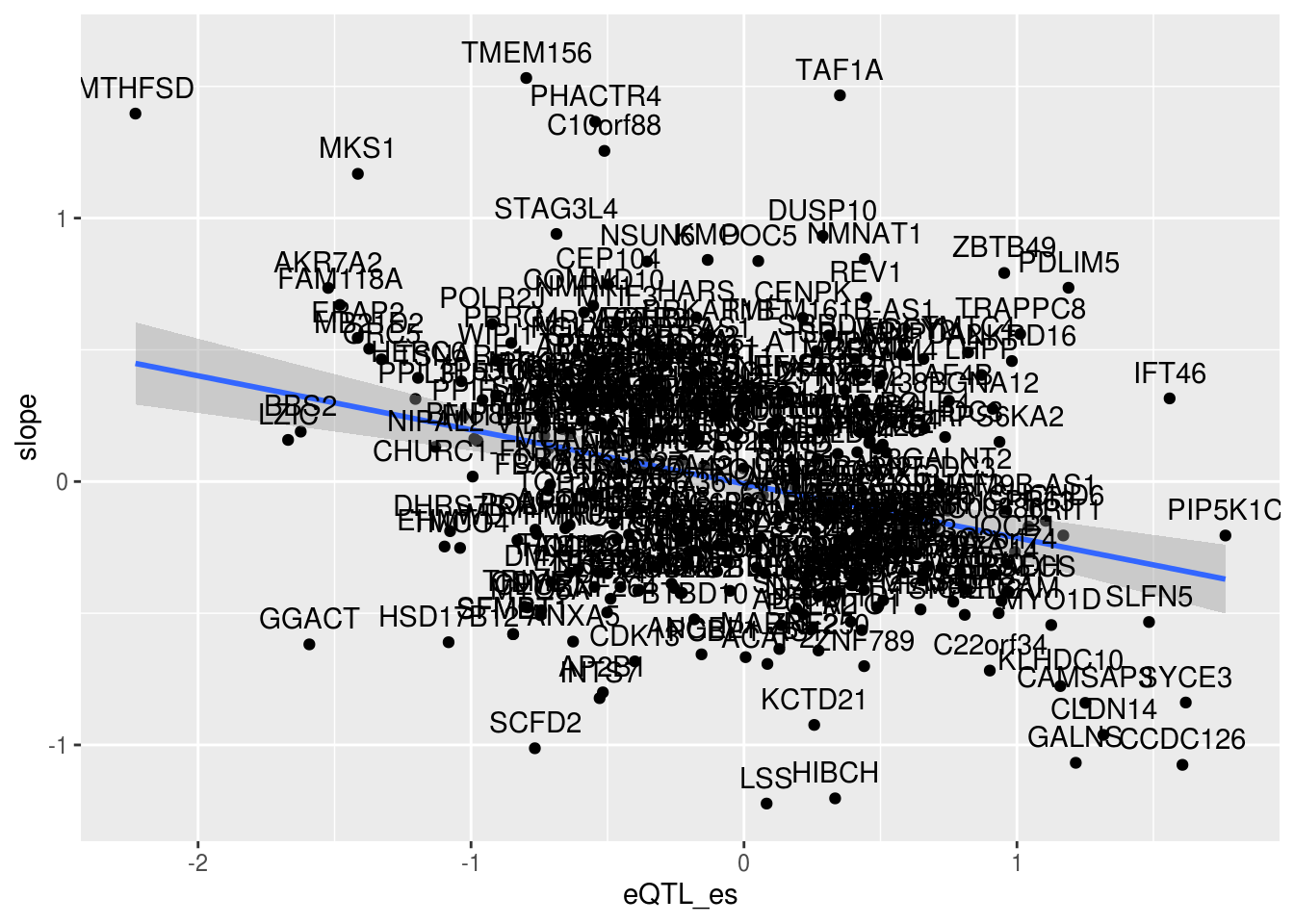 Number of PAS per gene:
Number of PAS per gene:
PAS=read.table("../data/PAS/APApeak_Peaks_GeneLocAnno.Nuclear.5perc.bed",col.names = c("chr", "start", "end", "name", "score", "strand"), stringsAsFactors = F) %>% separate(name, into=c('num', 'gene', 'loc'),sep=":" )
PASscore= PAS %>% select(num, score) %>% mutate(PASnum=paste("peak", num, sep=""))
PASgene=PAS %>% group_by(gene) %>% summarise(nPAS=n())alleQTLS_nuclear_genenum= alleQTLS_nuclear %>% inner_join(PASgene, by="gene")
alleQTLS_nuclear_score= alleQTLS_nuclear %>% inner_join(PASscore, by="PASnum")ggplot(alleQTLS_nuclear_genenum,aes(x=eQTL_es, y=slope,col=nPAS)) + geom_point() + geom_smooth(method = "lm") + geom_text(aes(label=gene), nudge_y = .1) 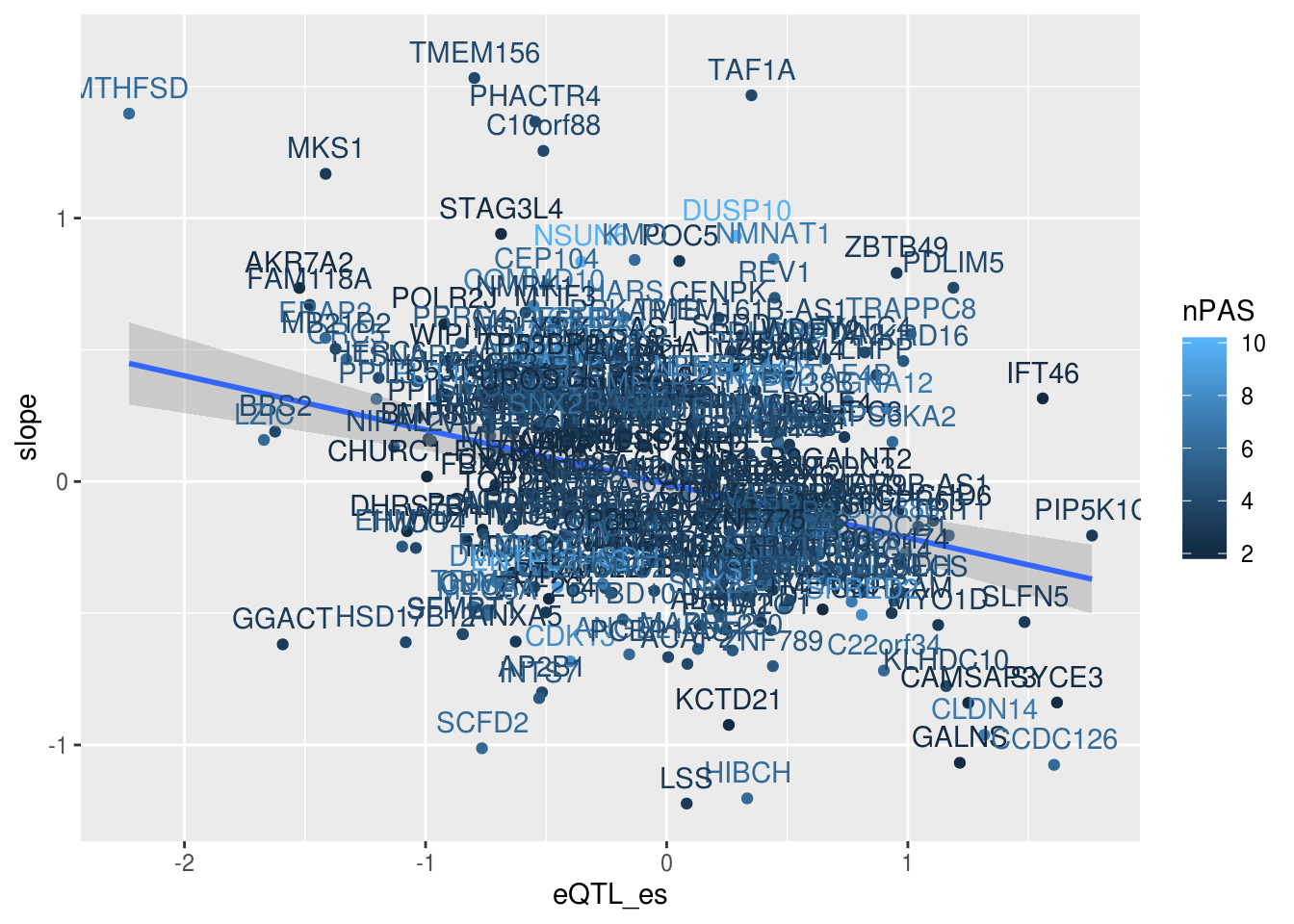
ggplot(alleQTLS_nuclear_score,aes(x=eQTL_es, y=slope,col=score)) + geom_point() + geom_smooth(method = "lm") + geom_text(aes(label=gene), nudge_y = .1) 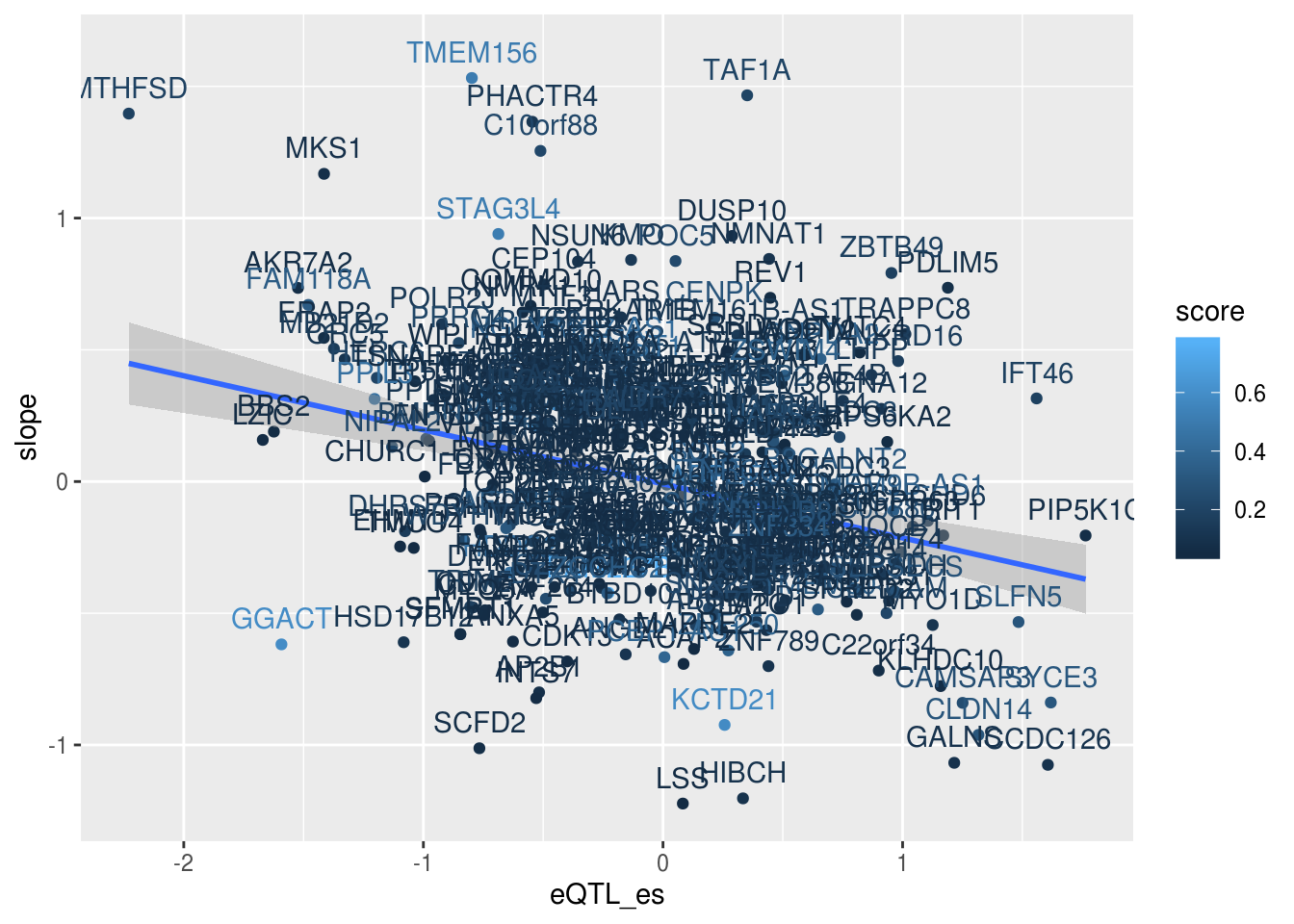
PAS %>% filter(gene=="MTHFSD") chr start end num gene loc score strand
1 16 86563782 86563783 52507 MTHFSD utr3 0.16923077 -
2 16 86564455 86564456 52509 MTHFSD utr3 0.31730769 -
3 16 86566995 86566996 52510 MTHFSD intron 0.18134615 -
4 16 86572985 86572986 52511 MTHFSD intron 0.05865385 -
5 16 86586779 86586780 52514 MTHFSD intron 0.05211538 -
6 16 86588585 86588586 52515 MTHFSD intron 0.20423077 -PAS %>% filter(gene=="TMEM156") chr start end num gene loc score strand
1 4 38967631 38967632 96732 TMEM156 end 0.10269231 -
2 4 38968368 38968369 96733 TMEM156 utr3 0.13307692 -
3 4 38985526 38985527 96738 TMEM156 intron 0.07557692 -
4 4 39029992 39029993 96746 TMEM156 intron 0.50500000 -plot expression and apa:
genohead=as.data.frame(read.table("../data/ExampleQTLPlots/genotypeHeader.txt", stringsAsFactors = F, header = F)[,10:128] %>% t()) %>% rename("Ind"=V1)
genotype=as.data.frame(read.table("../data/ExampleQTLPlots/TMEM156_NuclearPeaksGenotype.txt", stringsAsFactors = F, header = F) [,10:128] %>% t())ENSG00000121895 TMEM156
rs2711981
ref = c t=alt
RNAhead=as.data.frame(read.table("../data/molPhenos/RNAhead.txt", stringsAsFactors = F, header = F)[,5:73] %>% t())
RNApheno=as.data.frame(read.table("../data/molPhenos/RNA_TMEM156.txt", stringsAsFactors = F, header = F) [,5:73] %>% t())full_geno=bind_cols(Ind=genohead$Ind, dose=genotype$V1) %>% mutate(numdose=round(dose), genotype=ifelse(numdose==0, "CC", ifelse(numdose==1, "CT", "TT")))
full_pheno=bind_cols(Ind=RNAhead$V1, Expression=RNApheno$V1)
allRNA=full_geno %>% inner_join(full_pheno, by="Ind")Warning: Column `Ind` joining factors with different levels, coercing to
character vectorallRNA$genotype=as.factor(allRNA$genotype)
ggplot(allRNA, aes(x=genotype, y=Expression,group=genotype, fill=genotype)) + geom_boxplot() + geom_jitter()+scale_fill_brewer(palette = "Dark2") + labs(title="") + theme(legend.position = "bottom")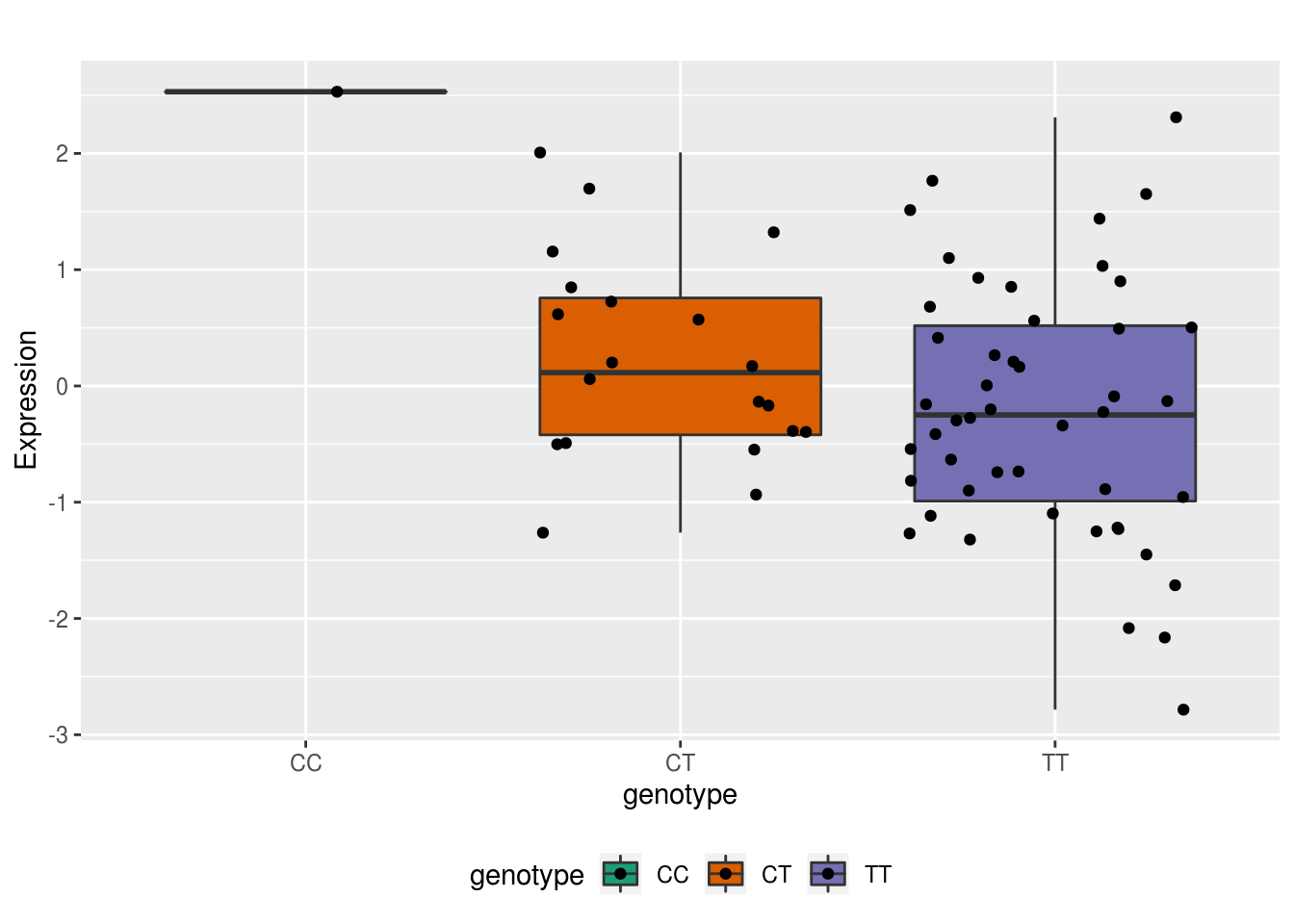
summary(lm(data=allRNA, Expression~genotype))
Call:
lm(formula = Expression ~ genotype, data = allRNA)
Residuals:
Min 1Q Median 3Q Max
-2.57048 -0.72008 -0.05794 0.70675 2.52445
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.531 1.044 2.423 0.0181 *
genotypeCT -2.303 1.070 -2.152 0.0350 *
genotypeTT -2.744 1.055 -2.601 0.0115 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.044 on 66 degrees of freedom
Multiple R-squared: 0.1164, Adjusted R-squared: 0.08964
F-statistic: 4.348 on 2 and 66 DF, p-value: 0.01683KCTD21 11 rs138463614 peak24232
ENSG00000188997
C is ref T is alt
genotype=as.data.frame(read.table("../data/ExampleQTLPlots/KCTD21_NuclearPeaksGenotype.txt", stringsAsFactors = F, header = F) [,10:128] %>% t())
RNApheno=as.data.frame(read.table("../data/molPhenos/RNA_KCTD21.txt", stringsAsFactors = F, header = F) [,5:73] %>% t())
full_geno=bind_cols(Ind=genohead$Ind, dose=genotype$V1) %>% mutate(numdose=round(dose), genotype=ifelse(numdose==0, "CC", ifelse(numdose==1, "CT", "TT")))
full_pheno=bind_cols(Ind=RNAhead$V1, Expression=RNApheno$V1)
allRNA=full_geno %>% inner_join(full_pheno, by="Ind")Warning: Column `Ind` joining factors with different levels, coercing to
character vectorallRNA$genotype=as.factor(allRNA$genotype)
ggplot(allRNA, aes(x=genotype, y=Expression,group=genotype, fill=genotype)) + geom_boxplot() + geom_jitter()+scale_fill_brewer(palette = "Dark2") + labs(title="") + theme(legend.position = "bottom")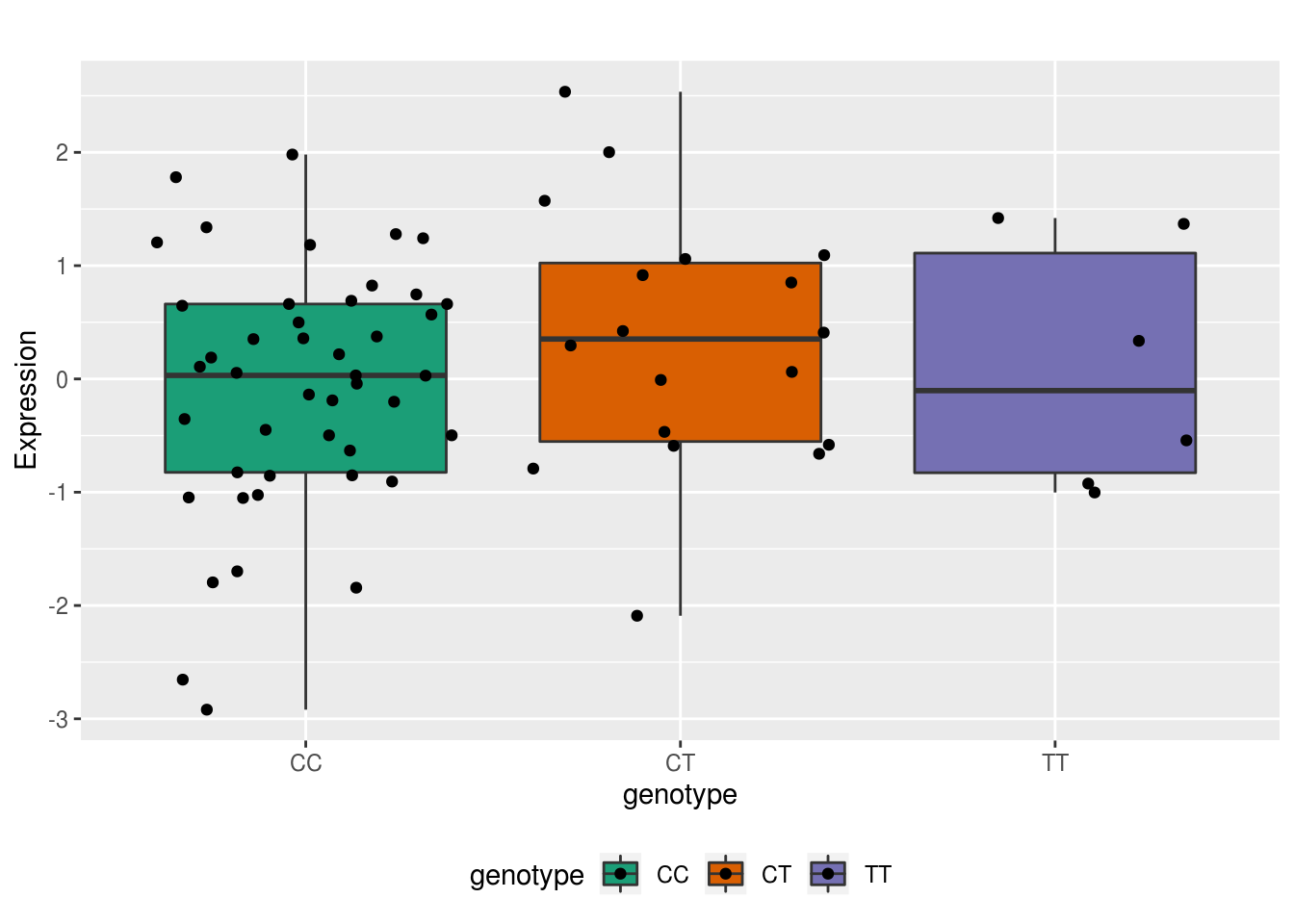
summary(lm(data=allRNA, Expression~genotype))
Call:
lm(formula = Expression ~ genotype, data = allRNA)
Residuals:
Min 1Q Median 3Q Max
-2.84260 -0.77776 0.08783 0.73800 2.20026
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.07669 0.16303 -0.470 0.640
genotypeCT 0.41155 0.30500 1.349 0.182
genotypeTT 0.18620 0.47530 0.392 0.697
Residual standard error: 1.094 on 66 degrees of freedom
Multiple R-squared: 0.02716, Adjusted R-squared: -0.002324
F-statistic: 0.9212 on 2 and 66 DF, p-value: 0.4031not a great example. expression effect size isnt strong.. but the diection is in the correct direction. increased expression with T.
filter to find another example:
alleQTLS_nuclear_scorefilt= alleQTLS_nuclear_score %>% filter(eQTL_es <1)
ggplot(alleQTLS_nuclear_scorefilt,aes(x=eQTL_es, y=slope,col=score)) + geom_point() + geom_text(aes(label=gene), nudge_y = .1) 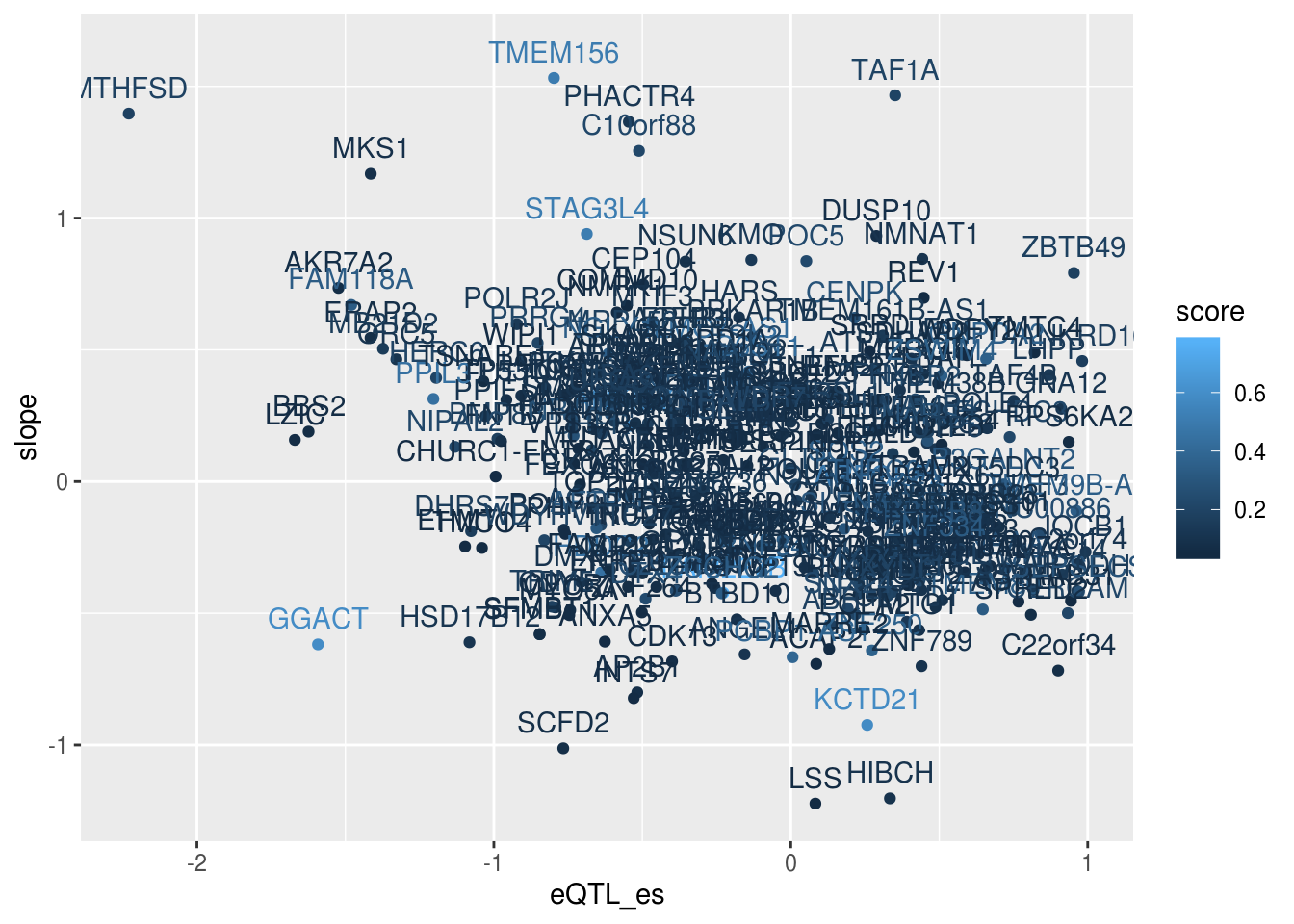
C10orf88 (ex in paper) peak 19682
rs7904973
ref =g alt = t
ENSG00000119965
genotype=as.data.frame(read.table("../data/ExampleQTLPlots/KCTD21_NuclearPeaksGenotype.txt", stringsAsFactors = F, header = F) [,10:128] %>% t())
RNApheno=as.data.frame(read.table("../data/molPhenos/RNA_c10orf88.txt", stringsAsFactors = F, header = F) [,5:73] %>% t())
full_geno=bind_cols(Ind=genohead$Ind, dose=genotype$V1) %>% mutate(numdose=round(dose), genotype=ifelse(numdose==0, "GG", ifelse(numdose==1, "GT", "TT")))
full_pheno=bind_cols(Ind=RNAhead$V1, Expression=RNApheno$V1)
allRNA=full_geno %>% inner_join(full_pheno, by="Ind")Warning: Column `Ind` joining factors with different levels, coercing to
character vectorallRNA$genotype=as.factor(allRNA$genotype)
ggplot(allRNA, aes(x=genotype, y=Expression,group=genotype, fill=genotype)) + geom_boxplot() + geom_jitter()+scale_fill_brewer(palette = "Dark2") + labs(title="") + theme(legend.position = "bottom")
summary(lm(data=allRNA, Expression~genotype))
Call:
lm(formula = Expression ~ genotype, data = allRNA)
Residuals:
Min 1Q Median 3Q Max
-2.86310 -0.52048 0.09394 0.52691 2.42987
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.10807 0.15391 0.702 0.485
genotypeGT -0.36784 0.28794 -1.277 0.206
genotypeTT -0.07503 0.44873 -0.167 0.868
Residual standard error: 1.032 on 66 degrees of freedom
Multiple R-squared: 0.0242, Adjusted R-squared: -0.005373
F-statistic: 0.8183 on 2 and 66 DF, p-value: 0.4456same situation with not a great effect size effect but again we see the correct direction.
sessionInfo()R version 3.6.1 (2019-07-05)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Scientific Linux 7.4 (Nitrogen)
Matrix products: default
BLAS/LAPACK: /software/openblas-0.2.19-el7-x86_64/lib/libopenblas_haswellp-r0.2.19.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] forcats_0.4.0 stringr_1.4.0 dplyr_0.8.3 purrr_0.3.2
[5] readr_1.3.1 tidyr_0.8.3 tibble_2.1.3 ggplot2_3.2.1
[9] tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] tidyselect_0.2.5 xfun_0.8 haven_2.1.1
[4] lattice_0.20-38 colorspace_1.4-1 generics_0.0.2
[7] vctrs_0.2.0 htmltools_0.3.6 yaml_2.2.0
[10] rlang_0.4.0 later_0.8.0 pillar_1.4.2
[13] withr_2.1.2 glue_1.3.1 RColorBrewer_1.1-2
[16] modelr_0.1.4 readxl_1.3.1 lifecycle_0.1.0
[19] munsell_0.5.0 gtable_0.3.0 workflowr_1.6.0
[22] cellranger_1.1.0 rvest_0.3.4 evaluate_0.14
[25] labeling_0.3 knitr_1.23 httpuv_1.5.1
[28] broom_0.5.2 Rcpp_1.0.3 promises_1.0.1
[31] backports_1.1.4 scales_1.1.0 jsonlite_1.6
[34] farver_2.0.1 fs_1.3.1 hms_0.5.0
[37] digest_0.6.20 stringi_1.4.3 grid_3.6.1
[40] rprojroot_1.3-2 cli_1.1.0 tools_3.6.1
[43] magrittr_1.5 lazyeval_0.2.2 crayon_1.3.4
[46] whisker_0.3-2 pkgconfig_2.0.2 zeallot_0.1.0
[49] xml2_1.2.0 lubridate_1.7.4 assertthat_0.2.1
[52] rmarkdown_1.13 httr_1.4.0 rstudioapi_0.10
[55] R6_2.4.0 nlme_3.1-140 git2r_0.26.1
[58] compiler_3.6.1