apaQTL GWAS overlap
Briana Mittleman
10/26/2018
Last updated: 2018-11-15
workflowr checks: (Click a bullet for more information)-
✔ R Markdown file: up-to-date
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
-
✔ Environment: empty
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
-
✔ Seed:
set.seed(12345)The command
set.seed(12345)was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible. -
✔ Session information: recorded
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
-
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.✔ Repository version: 7960cbb
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can usewflow_publishorwflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.Ignored files: Ignored: .DS_Store Ignored: .Rhistory Ignored: .Rproj.user/ Ignored: data/.DS_Store Ignored: output/.DS_Store Untracked files: Untracked: KalistoAbundance18486.txt Untracked: analysis/ncbiRefSeq_sm.sort.mRNA.bed Untracked: analysis/snake.config.notes.Rmd Untracked: analysis/verifyBAM.Rmd Untracked: data/18486.genecov.txt Untracked: data/APApeaksYL.total.inbrain.bed Untracked: data/ChromHmmOverlap/ Untracked: data/GM12878.chromHMM.bed Untracked: data/GM12878.chromHMM.txt Untracked: data/LocusZoom/ Untracked: data/NuclearApaQTLs.txt Untracked: data/PeaksUsed/ Untracked: data/RNAkalisto/ Untracked: data/TotalApaQTLs.txt Untracked: data/Totalpeaks_filtered_clean.bed Untracked: data/YL-SP-18486-T-combined-genecov.txt Untracked: data/YL-SP-18486-T_S9_R1_001-genecov.txt Untracked: data/apaExamp/ Untracked: data/bedgraph_peaks/ Untracked: data/bin200.5.T.nuccov.bed Untracked: data/bin200.Anuccov.bed Untracked: data/bin200.nuccov.bed Untracked: data/clean_peaks/ Untracked: data/comb_map_stats.csv Untracked: data/comb_map_stats.xlsx Untracked: data/comb_map_stats_39ind.csv Untracked: data/combined_reads_mapped_three_prime_seq.csv Untracked: data/diff_iso_trans/ Untracked: data/ensemble_to_genename.txt Untracked: data/example_gene_peakQuant/ Untracked: data/filtered_APApeaks_merged_allchrom_refseqTrans.closest2End.bed Untracked: data/filtered_APApeaks_merged_allchrom_refseqTrans.closest2End.noties.bed Untracked: data/first50lines_closest.txt Untracked: data/gencov.test.csv Untracked: data/gencov.test.txt Untracked: data/gencov_zero.test.csv Untracked: data/gencov_zero.test.txt Untracked: data/gene_cov/ Untracked: data/joined Untracked: data/leafcutter/ Untracked: data/merged_combined_YL-SP-threeprimeseq.bg Untracked: data/mol_overlap/ Untracked: data/mol_pheno/ Untracked: data/nom_QTL/ Untracked: data/nom_QTL_opp/ Untracked: data/nom_QTL_trans/ Untracked: data/nuc6up/ Untracked: data/other_qtls/ Untracked: data/peakPerRefSeqGene/ Untracked: data/perm_QTL/ Untracked: data/perm_QTL_opp/ Untracked: data/perm_QTL_trans/ Untracked: data/reads_mapped_three_prime_seq.csv Untracked: data/smash.cov.results.bed Untracked: data/smash.cov.results.csv Untracked: data/smash.cov.results.txt Untracked: data/smash_testregion/ Untracked: data/ssFC200.cov.bed Untracked: data/temp.file1 Untracked: data/temp.file2 Untracked: data/temp.gencov.test.txt Untracked: data/temp.gencov_zero.test.txt Untracked: output/picard/ Untracked: output/plots/ Untracked: output/qual.fig2.pdf Unstaged changes: Modified: analysis/28ind.peak.explore.Rmd Modified: analysis/39indQC.Rmd Modified: analysis/cleanupdtseq.internalpriming.Rmd Modified: analysis/coloc_apaQTLs_protQTLs.Rmd Modified: analysis/dif.iso.usage.leafcutter.Rmd Modified: analysis/diff_iso_pipeline.Rmd Modified: analysis/explore.filters.Rmd Modified: analysis/flash2mash.Rmd Modified: analysis/overlapMolQTL.Rmd Modified: analysis/overlap_qtls.Rmd Modified: analysis/peakOverlap_oppstrand.Rmd Modified: analysis/pheno.leaf.comb.Rmd Modified: analysis/swarmPlots_QTLs.Rmd Modified: analysis/test.max2.Rmd Modified: code/Snakefile
Expand here to see past versions:
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 7960cbb | Briana Mittleman | 2018-11-15 | new plink call |
| html | 4a4b5c7 | Briana Mittleman | 2018-11-12 | Build site. |
| Rmd | 89b780f | Briana Mittleman | 2018-11-12 | add analysis of hits |
| html | 7462a2d | Briana Mittleman | 2018-11-09 | Build site. |
| Rmd | 489bf91 | Briana Mittleman | 2018-11-09 | code for plink r2 |
| html | 0428c8c | Briana Mittleman | 2018-10-29 | Build site. |
| Rmd | 42d0e3d | Briana Mittleman | 2018-10-29 | add gwas overlap to index |
library(workflowr)This is workflowr version 1.1.1
Run ?workflowr for help getting startedlibrary(tidyverse)── Attaching packages ─────────────────────────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.0.0 ✔ purrr 0.2.5
✔ tibble 1.4.2 ✔ dplyr 0.7.6
✔ tidyr 0.8.1 ✔ stringr 1.3.1
✔ readr 1.1.1 ✔ forcats 0.3.0── Conflicts ────────────────────────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()library(cowplot)
Attaching package: 'cowplot'The following object is masked from 'package:ggplot2':
ggsaveIn this analysis I want to see if APAqtls show up in the GWAS catelog. I then want to see if they explain different signal then overlappnig the eQTLs.
I can use my significant snp bed file from /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps to overlap with the GWAS catelog. First I can look at direct location then I will use an LD cutoff to colocalize.
- /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Nuclear.sort.bed
- /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Total.sort.bed
The downloaded GWAS catalog from the UCSD table browser.
- /project2/gilad/briana/genome_anotation_data/hg19GwasCatalog.txt
I will make this into a bed format to use with pybedtools.
-Chrom -start -end -name -score
fin=open(""/project2/gilad/briana/genome_anotation_data/hg19GwasCatalog.txt", "r")
fout=open("/project2/gilad/briana/genome_anotation_data/hg19GwasCatalog.bed","w")
for num, ln in enumerate(fin):
if num > 0:
line=ln.split("\t")
id_list=[line[4],line[5], line[14]]
start=int(line[2])
end=int(line[3])
id=":".join(id_list)
chr=line[1][3:]
pval=line[16]
fout.write("%s\t%d\t%d\t%s\t%s\n"%(chr,start, end, id, pval)
fout.close()
Pybedtools to intersect my snps with catelog /project2/gilad/briana/threeprimeseq/data/GWAS_overlap
output dir:
import pybedtools
gwas=pybedtools.BedTool("/project2/gilad/briana/genome_anotation_data/hg19GwasCatalog.sort.bed")
nuc=pybedtools.BedTool("/project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Nuclear.sort.bed")
tot=pybedtools.BedTool("/project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Total.sort.bed")
nucOverGWAS=nuc.intersect(gwas, wa=True,wb=True)
totOverGWAS=tot.intersect(gwas,wa=True, wb=True)
#this only results in one overlap:
nucOverGWAS.saveas("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/nucFDR10overlapGWAS.txt")
Problem: I see this snp but it is assoicated with a different gene. I need to think about gene and snp overlap.
I can see if this snp is an eqtl.
16:30482494
eqtl=read.table(file = "../data/other_qtls/fastqtl_qqnorm_RNAseq_phase2.fixed.perm.out")
eqtl_g= read.table("../data/other_qtls/fastqtl_qqnorm_RNAseqGeuvadis.fixed.perm.out")This snp is not in either of these files. I will check for them in the nominal results.
grep 16:30482494 /project2/gilad/briana/threeprimeseq/data/molecular_QTLs/nom/fastqtl_qqnorm_RNAseq_phase2.fixed.nominal.out
grep 16:30482494 /project2/gilad/briana/threeprimeseq/data/molecular_QTLs/nom/fastqtl_qqnorm_RNAseqGeuvadis.fixed.nominal.out
LD structure
https://vcftools.github.io/man_latest.html –vcf (vcf file) –geno-r2 –out (prefix) vcf tools is on midway 2 “module load vcftools”
I can use the snp files I created for the chromHMM analysis.
- /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Total.sort.bed
- /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnp/ApaQTLsignificantSnps_10percFDR_Nuclear.sort.bed
I can use awk to get the first and third column.
awk '{print $1 ":" $3}' /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Nuclear.sort.bed > /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear.txt
awk '{print $1":"$3}' /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/sigSnps/ApaQTLsignificantSnps_10percFDR_Total.sort.bed > /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txttestLD_vcftools_totQTL.sh
#!/bin/bash
#SBATCH --job-name=testLD_vcftools_totQTL.sh
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=testLD_vcftools_totQTL.out
#SBATCH --error=testLD_vcftools_totQTL.err
#SBATCH --partition=broadwl
#SBATCH --mem=16G
#SBATCH --mail-type=END
module load vcftools
vcftools --gzvcf chr1.dose.vcf.gz --snps /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt --out /project2/gilad/briana/YRI_geno_hg19/chr1.totQTL.LD --geno-r2 /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD
/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD
Now run this for all chr in both fractions.
LD_vcftools.sh
#!/bin/bash
#SBATCH --job-name=LD_vcftools.sh
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=LD_vcftools.out
#SBATCH --error=rLD_vcftools.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load vcftools
for i in {1..22};
do
vcftools --gzvcf /project2/gilad/briana/YRI_geno_hg19/chr${i}.dose.vcf.gz --snps /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD/chr${i}.totQTL.LD --geno-r2 --min-r2 .8
done
for i in {1..22};
do
vcftools --gzvcf /project2/gilad/briana/YRI_geno_hg19/chr${i}.dose.vcf.gz --snps /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear.txt --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD/chr${i}.nucQTL.LD --geno-r2 --min-r2 .8
done
This doesnt give very many more snps. Let me try this with Tony’s vcf files from the larger panel of LCLs.
Try it with the –hap-r2 argument.
LD_vcftools.hap.sh
#!/bin/bash
#SBATCH --job-name=LD_vcftools.hap.sh
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=LD_vcftools.hap.out
#SBATCH --error=rLD_vcftools.hap.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load vcftools
for i in {1..22};
do
vcftools --gzvcf /project2/gilad/briana/YRI_geno_hg19/chr${i}.dose.vcf.gz --snps /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD/chr${i}.totQTL.hap.LD --hap-r2--min-r2 .8
done
for i in {1..22};
do
vcftools --gzvcf /project2/gilad/briana/YRI_geno_hg19/chr${i}.dose.vcf.gz --snps /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear.txt --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD/chr${i}.nucQTL.hap.LD --hap-r2 --min-r2 .8
done
still not a lot of snps.
testLDGeu_vcftools_totQTL.sh
#!/bin/bash
#SBATCH --job-name=testLDGeu_vcftools_totQTL.sh
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=testLDGeu_vcftools_totQTL.out
#SBATCH --error=testLDGeu_vcftools_totQTL.err
#SBATCH --partition=broadwl
#SBATCH --mem=16G
#SBATCH --mail-type=END
module load vcftools
vcftools --gzvcf /project2/yangili1/LCL/genotypesYRI.gen.txt.gz --snps /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/geuvadis.totQTL.LD --geno-r2 Error: Insufficient sites remained after filtering
vcf2Plink.sh
#!/bin/bash
#SBATCH --job-name=vcf2Plink
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=vcf2Plink.out
#SBATCH --error=vcf2Plink.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load vcftools
for i in {1..22};
do
vcftools --gzvcf /project2/gilad/briana/YRI_geno_hg19/chr${i}.dose.vcf.gz --plink --chr ${i} --out /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chr${i}
doneTry with plink:
I will use the ped and map files: –ped /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chr$i.ped –map /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chri.map
–ld-snp-list /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt
–r2
–ld-window-r2 0.20.8 testPlink_r2.sh
#!/bin/bash
#SBATCH --job-name=testPlink_r2
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=testPlink_r2.out
#SBATCH --error=testPlink_r2.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load plink
plink --ped /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chr22.ped --map /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chr22.map --r2 --ld-window-r2 0.8 --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/plinkYRI_LDchr22
This gives me 77,000 pairs. I will run this on all of the chromosomes then subset by snps i have QTLs for.
RunPlink_r2.sh
#!/bin/bash
#SBATCH --job-name=RunPlink_r2
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=RunPlink_r2.out
#SBATCH --error=RunPlink_r2.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load plink
for i in {1..22};
do
plink --ped /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chr${i}.ped --map /project2/gilad/briana/YRI_geno_hg19/plinkYRIgeno_chr${i}.map --r2 --ld-window-r2 0.8 --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/plinkYRI_LDchr${i}
done
I can now subset these files for snps in the /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt and /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear.txt files using a python script.
This script will take a fraction and chromosome.
subset_plink4QTLs.py
def main(genFile, qtlFile, outFile):
#convert snp file to a list:
def file_to_list(file):
snp_list=[]
for ln in file:
snp=ln.strip()
snp_list.append(snp)
return(snp_list)
gen=open(genFile,"r")
fout=open(outFile, "w")
qtls=open(qtlFile, "r")
qtl_list=file_to_list(qtls)
for ln in gen:
snp=ln.split()[2]
if snp in qtl_list:
fout.write(ln)
fout.close()
if __name__ == "__main__":
import sys
chrom=sys.argv[1]
fraction=sys.argv[2]
genFile = "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/plinkYRI_LDchr%s.ld"%(chrom)
outFile= "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/%sApaQTL_LD/chr%s.%sQTL.LD.geno.ld"%(fraction,chrom,fraction)
qtlFile= "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_%s.txt"%(fraction)
main(genFile, qtlFile, outFile)
Run this for all chr in a bash script:
run_subset_plink4QTLs.sh
#!/bin/bash
#SBATCH --job-name=run_subset_plink4QTLs
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=run_subset_plink4QTLs.out
#SBATCH --error=run_subset_plink4QTLs.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load Anaconda3
source activate three-prime-env
for i in {1..22};
do
python subset_plink4QTLs.py ${i} "Total"
done
for i in {1..22};
do
python subset_plink4QTLs.py ${i} "Nuclear"
doneThis results in 385 more snps for the nuclear QTLs and 54 more for the total.
I want to try this method on the bigger panel from Tonys work.
vcf2Plink_geu.sh
#!/bin/bash
#SBATCH --job-name=vcf2Plink_geu
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=vcf2Plink_geu2.out
#SBATCH --error=vcf2Plink_geu2.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load vcftools
for i in {1..22};
do
vcftools --gzvcf /project2/yangili1/LCL/geuvadis_genotypes/GEUVADIS.chr${i}.hg19_MAF5AC.vcf.gz --plink --chr ${i} --out /project2/gilad/briana/YRI_geno_hg19/geu_plinkYRIgeno_chr${i}
doneRunPlink_Geu_r2.sh
#!/bin/bash
#SBATCH --job-name=RunPlink_geu_r2
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=RunPlink_geu_r2.out
#SBATCH --error=RunPlink_geu_r2.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load plink
for i in {1..22};
do
plink --ped /project2/gilad/briana/YRI_geno_hg19/geu_plinkYRIgeno_chr${i}.ped --map /project2/gilad/briana/YRI_geno_hg19/geu_plinkYRIgeno_chr${i}.map --r2 --ld-window-r2 0.8 --out /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/geu_plinkYRI_LDchr${i}
doneQTLs2GeuSnps.py
tot_in=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total.txt", "r")
nuc_in=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear.txt", "r")
tot_out=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total_GEU.txt", "w")
nuc_out=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear_GEU.txt", "w")
def fix_file(fin, fout):
for ln in fin:
chrom, pos = ln.split(":")
fout.write("snp_%s_%s/n"%(chrom,pos))
fout.close()
fix_file(tot_in, tot_out)
fix_file(nuc_in, nuc_out)
run_QTLs2GeuSnps.sh
#!/bin/bash
#SBATCH --job-name=run_QTLs2GeuSnps
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=run_QTLs2GeuSnps.out
#SBATCH --error=run_QTLs2GeuSnps.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load Anaconda3
source activate three-prime-env
python QTLs2GeuSnps.pyUpdate the python selection script for geu results.
subset_plink4QTLs_geu.py
def main(genFile, qtlFile, outFile):
#convert snp file to a list:
def file_to_list(file):
snp_list=[]
for ln in file:
snp=ln.strip()
snp_list.append(snp)
return(snp_list)
gen=open(genFile,"r")
fout=open(outFile, "w")
qtls=open(qtlFile, "r")
qtl_list=file_to_list(qtls)
for ln in gen:
snp=ln.split()[2]
if snp in qtl_list:
fout.write(ln)
fout.close()
if __name__ == "__main__":
import sys
chrom=sys.argv[1]
fraction=sys.argv[2]
genFile = "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/geu_plinkYRI_LDchr%s.ld"%(chrom)
outFile= "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/%sApaQTL_LD_geu/chr%s.%sQTL.LD.geno.ld"%(fraction,chrom,fraction)
qtlFile= "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_%s_GEU.txt"%(fraction)
main(genFile, qtlFile, outFile)
run_subset_plink4QTLs_geu.sh
#!/bin/bash
#SBATCH --job-name=run_subset_plink4QTLs_geu
#SBATCH --account=pi-yangili1
#SBATCH --time=36:00:00
#SBATCH --output=run_subset_plink4QTLs_geu.out
#SBATCH --error=run_subset_plink4QTLs_geu.err
#SBATCH --partition=broadwl
#SBATCH --mem=30G
#SBATCH --mail-type=END
module load Anaconda3
source activate three-prime-env
for i in {1..22};
do
python subset_plink4QTLs_geu.py ${i} "Total"
done
for i in {1..22};
do
python subset_plink4QTLs_geu.py ${i} "Nuclear"
doneThis add 1228 for total and 10251 for nuclear. This is better. I will use these for the GWAS overlap.
I want to make a sorted bed file with all of these snps (total and nuclear together) to overlap with the gwas catelog. I will have the snp name include if it was a from the total or nuclear. I can do all of this in python then sort the bed file after.
The LD files include indels. I will not include there. There are 8 in the total file and 108 in nuclear, I can remove these with the following.
grep -v indel /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD_geu/allChr.NuclearQTL.LD.gene.ld > /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD_geu/allChr.NuclearQTL.LD.gene.ld_noIndel
grep -v indel /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD_geu/allChr.TotalQTL.GD.geno.ld > /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD_geu/allChr.TotalQTL.GD.geno.ld_noIndel- /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total_GEU.txt
- /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear_GEU.txt
- /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD_geu/allChr.NuclearQTL.LD.gene.ld_noIndel
- /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD_geu/allChr.TotalQTL.GD.geno.ld_noIndel
makeAlloverlapbed.py
#load files:
QTL_total=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Total_GEU.txt", "r")
QTL_nuclear=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/ApaQTLsigSNPpos_Nuclear_GEU.txt", "r")
LD_total=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/TotalApaQTL_LD_geu/allChr.TotalQTL.GD.geno.ld_noIndel", "r")
LD_nuclear=open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD_geu/allChr.NuclearQTL.LD.gene.ld_noIndel", "r")
outFile= open("/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/AllOverlapSnps.bed", "w")
#function for qtl to bed format
def qtl2bed(fqtl, fraction, fout=outFile):
for ln in fqtl:
snp, chrom, pos = ln.split("_")
start=int(pos)-1
end= int(pos)
fout.write("%s\t%d\t%d\tQTL_%s\n"%(chrom, start, end,fraction))
#function for ld to bed format
def ld2bed(fLD, fraction, fout=outFile):
for ln in fLD:
snpID=ln.split()[5]
snp, chrom, pos= snpID.split("_")
start=int(pos)-1
end=int(pos)
fout.write("%s\t%d\t%d\tLD_%s\n"%(chrom, start, end,fraction))
#I will run each of these for both fractions to get all of the snps in the out file.
qtl2bed(QTL_nuclear, "Nuclear")
qtl2bed(QTL_total, "Total")
ld2bed(LD_nuclear, "Nuclear")
ld2bed(LD_total, "Total")
outFile.close()Sort it:
sort -k1,1 -k2,2n /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/AllOverlapSnps.bed > /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/AllOverlapSnps_sort.bedI can now use py bedtools to overlap this.
overlapSNPsGWAS.py
This will take in any lsit of snps and overlap them with the gwas catelog bed file.
def main(infile, outfile):
gwas_file=open("/project2/gilad/briana/genome_anotation_data/hg19GwasCatalog.sort.bed","r")
gwas=pybedtools.BedTool(gwas_file)
snps_file=open(infile, "r")
snps=pybedtools.BedTool(snps_file)
snpOverGWAS=snps.intersect(gwas, wa=True,wb=True)
snpOverGWAS.saveas(outfile)
if __name__ == "__main__":
import sys
import pybedtools
infile=sys.argv[1]
outfile=sys.argv[2]
main(infile, outfile) Call this in bash so i can load the environment
run_overlapSNPsGWAS.sh
#!/bin/bash
#SBATCH --job-name=run_overlapSNPsGWAS
#SBATCH --account=pi-yangili1
#SBATCH --time=5:00:00
#SBATCH --output=run_overlapSNPsGWAS.out
#SBATCH --error=run_overlapSNPsGWAS.err
#SBATCH --partition=broadwl
#SBATCH --mem=10G
#SBATCH --mail-type=END
module load Anaconda3
source activate three-prime-env
python overlapSNPsGWAS.py "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/AllOverlapSnps_sort.bed" "/project2/gilad/briana/threeprimeseq/data/GWAS_overlap/AllSnps_GWASoverlapped.txt"There are 13 overlaps now.
old stuff before change to plink
Still only get 2 that overlap the catelog. They are in the ITGAL and NCAPG genes. I should check if 16:30482494 (the nuclear QTL) is also a eQTL not how i did before but with my code from the all boxplot analysis
plotQTL_func= function(SNP, peak, gene){
apaN_file=read.table(paste("../data/apaExamp/qtlSNP_PeakAPANuclear.", SNP, peak, ".txt", sep = "" ), header=T)
apaT_file=read.table(paste("../data/apaExamp/qtlSNP_PeakAPATotal.", SNP, peak, ".txt", sep = "" ), header=T)
su30_file=read.table(paste("../data/apaExamp/qtlSNP_Peak_4su_30_", SNP, gene, ".txt", sep=""), header = T)
su60_file=read.table(paste("../data/apaExamp/qtlSNP_Peak_4su_60_", SNP, gene, ".txt", sep=""), header=T)
RNA_file=read.table(paste("../data/apaExamp/qtlSNP_Peak_RNAseq_", SNP, gene, ".txt", sep=""),header=T)
RNAg_file=read.table(paste("../data/apaExamp/qtlSNP_Peak_RNAseqGeuvadis_", SNP, gene, ".txt", sep=""), header = T)
ribo_file=read.table(paste("../data/apaExamp/qtlSNP_Peak_ribo_", SNP, gene, ".txt", sep=""),header=T)
prot_file=read.table(paste("../data/apaExamp/qtlSNP_Peak_prot.", SNP, gene, ".txt", sep=""), header=T)
ggplot_func= function(file, molPhen,GENE){
file = file %>% mutate(genotype=Allele1 + Allele2)
file$genotype= as.factor(as.character(file$genotype))
plot=ggplot(file, aes(y=Pheno, x=genotype, by=genotype, fill=genotype)) + geom_boxplot(width=.25) + geom_jitter() + labs(y="Phenotpye",title=paste(molPhen, GENE, sep=": ")) + scale_fill_brewer(palette="Paired")
return(plot)
}
apaNplot=ggplot_func(apaN_file, "Apa Nuclear", gene)
apaTplot=ggplot_func(apaT_file, "Apa Total", gene)
su30plot=ggplot_func(su30_file, "4su30",gene)
su60plot=ggplot_func(su60_file, "4su60",gene)
RNAplot=ggplot_func(RNA_file, "RNA Seq",gene)
RNAgPlot=ggplot_func(RNAg_file, "RNA Seq Geuvadis",gene)
riboPlot= ggplot_func(ribo_file, "Ribo Seq",gene)
protplot=ggplot_func(prot_file, "Protein",gene)
full_plot= plot_grid(apaNplot,apaTplot, su30plot, su60plot, RNAplot, RNAgPlot, riboPlot, protplot,nrow=2)
return (full_plot)
}16:30482494 PPP4C_+_peak122195
grep peak122195 /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Nuclear_transcript_permResBH.txt
#gene=PPP4C
grep PPP4C /project2/gilad/briana/genome_anotation_data/ensemble_to_genename.txt
#ensg= ENSG00000149923
python createQTLsnpAPAPhenTable.py 16 16:30482494 peak122195 Total
python createQTLsnpAPAPhenTable.py 16 16:30482494 peak122195 Nuclear
sbatch run_createQTLsnpMolPhenTable.sh "16" "16:30482494" "ENSG00000149923"
scp brimittleman@midway2.rcc.uchicago.edu:/project2/gilad/briana/threeprimeseq/data/ApaQTL_examples/*16:30482494* /Users/bmittleman1/Documents/Gilad_lab/threeprimeseq/data/apaExampplotQTL_func(SNP="16:30482494", peak="peak122195", gene="ENSG00000149923")Warning: Removed 2 rows containing non-finite values (stat_boxplot).Warning: Removed 2 rows containing missing values (geom_point).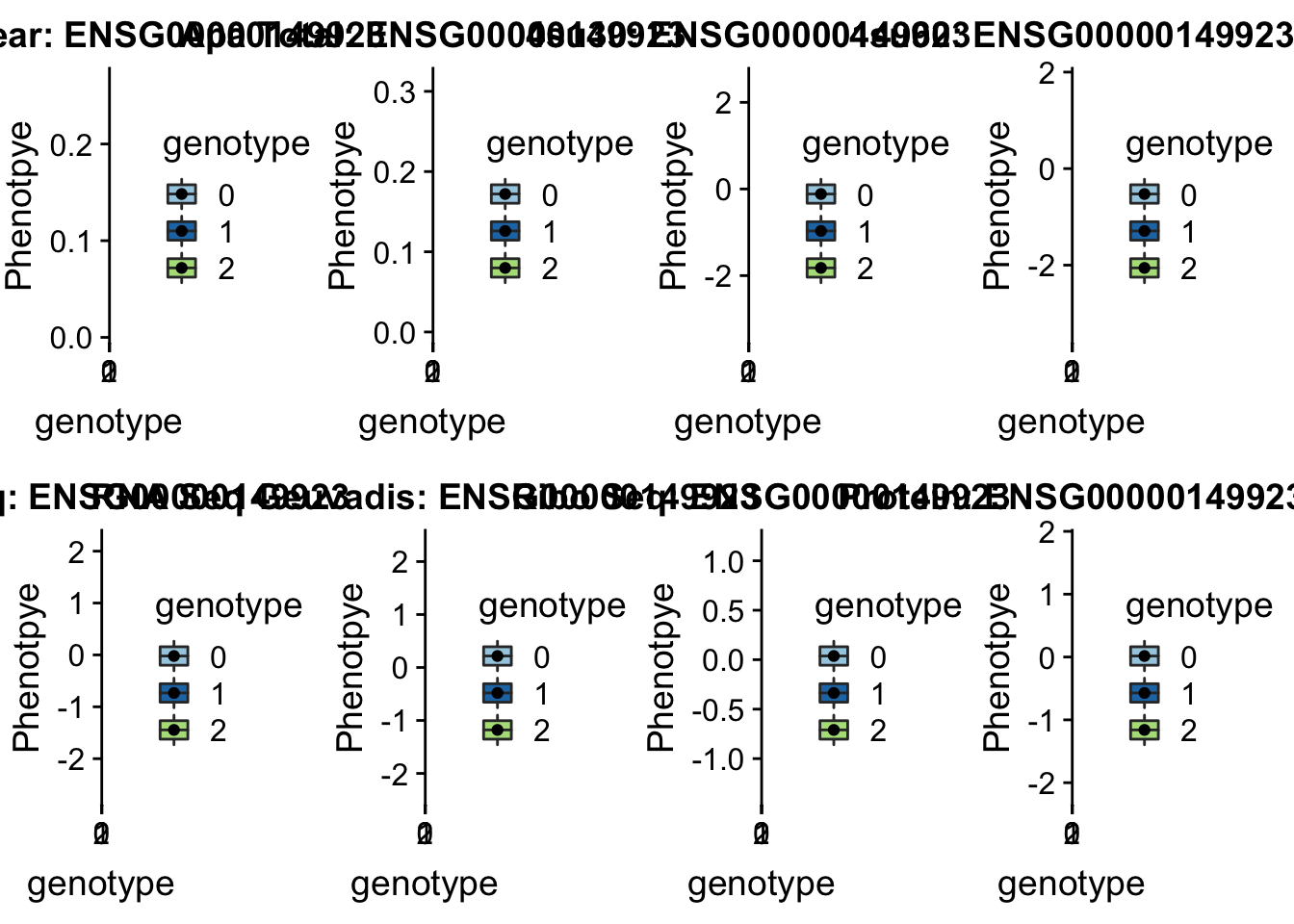
Expand here to see past versions of unnamed-chunk-29-1.png:
| Version | Author | Date |
|---|---|---|
| 4a4b5c7 | Briana Mittleman | 2018-11-12 |
This is in a GWAS for Ulcerative colitis.
I can look at the LD snp as well. I just need to check the ld snps and see which snp it corresponds to in my QTLs.
4:17797966
grep snp_4_17797966 /project2/gilad/briana/threeprimeseq/data/GWAS_overlap/NuclearApaQTL_LD_geu/allChr.NuclearQTL.LD.gene.ld_noIndel
In my analysis the snp is 4:17797455 DCAF16_-_peak236311: This is also a different gene.
grep peak236311 /project2/gilad/briana/threeprimeseq/data/perm_APAqtl_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Nuclear_transcript_permResBH.txt
#gene=PPP4C
grep DCAF16 /project2/gilad/briana/genome_anotation_data/ensemble_to_genename.txt
#ensg=ENSG00000163257
python createQTLsnpAPAPhenTable.py 4 4:17797455 peak236311 Total
python createQTLsnpAPAPhenTable.py 4 4:17797455 peak236311 Nuclear
sbatch run_createQTLsnpMolPhenTable.sh "4" "4:17797455" "ENSG00000163257"
scp brimittleman@midway2.rcc.uchicago.edu:/project2/gilad/briana/threeprimeseq/data/ApaQTL_examples/*4:17797455* /Users/bmittleman1/Documents/Gilad_lab/threeprimeseq/data/apaExampplotQTL_func(SNP="4:17797455", peak="peak236311", gene="ENSG00000163257")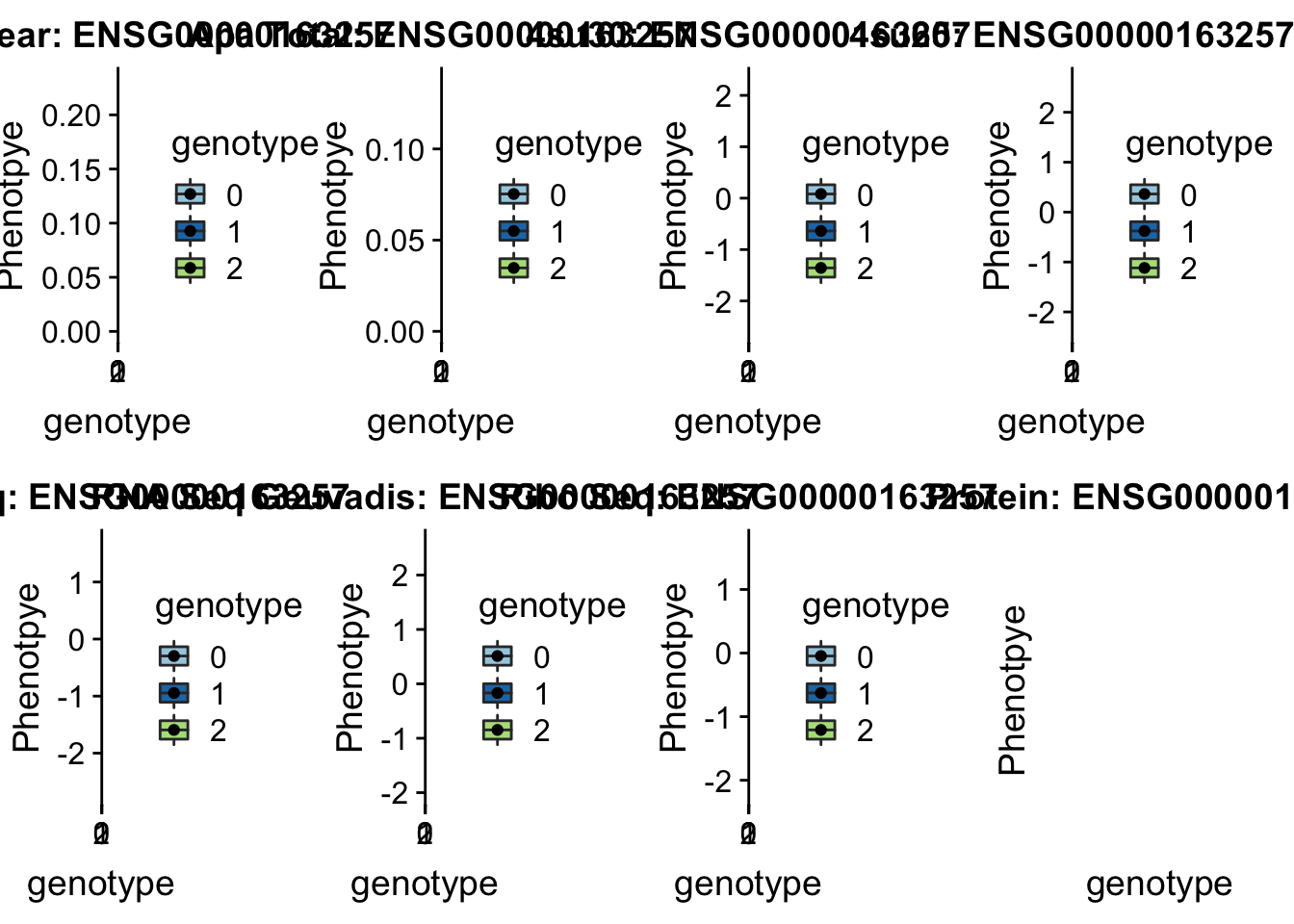
Expand here to see past versions of unnamed-chunk-32-1.png:
| Version | Author | Date |
|---|---|---|
| 4a4b5c7 | Briana Mittleman | 2018-11-12 |
This example is a GWAS hit for height .
Yangs script
–ld-window-kb 1000 –ld-window 99999 –ld-window-r2 0.8
Session information
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Sierra 10.12.6
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] bindrcpp_0.2.2 cowplot_0.9.3 forcats_0.3.0 stringr_1.3.1
[5] dplyr_0.7.6 purrr_0.2.5 readr_1.1.1 tidyr_0.8.1
[9] tibble_1.4.2 ggplot2_3.0.0 tidyverse_1.2.1 workflowr_1.1.1
loaded via a namespace (and not attached):
[1] tidyselect_0.2.4 haven_1.1.2 lattice_0.20-35
[4] colorspace_1.3-2 htmltools_0.3.6 yaml_2.2.0
[7] rlang_0.2.2 R.oo_1.22.0 pillar_1.3.0
[10] glue_1.3.0 withr_2.1.2 R.utils_2.7.0
[13] RColorBrewer_1.1-2 modelr_0.1.2 readxl_1.1.0
[16] bindr_0.1.1 plyr_1.8.4 munsell_0.5.0
[19] gtable_0.2.0 cellranger_1.1.0 rvest_0.3.2
[22] R.methodsS3_1.7.1 evaluate_0.11 labeling_0.3
[25] knitr_1.20 broom_0.5.0 Rcpp_0.12.19
[28] scales_1.0.0 backports_1.1.2 jsonlite_1.5
[31] hms_0.4.2 digest_0.6.17 stringi_1.2.4
[34] grid_3.5.1 rprojroot_1.3-2 cli_1.0.1
[37] tools_3.5.1 magrittr_1.5 lazyeval_0.2.1
[40] crayon_1.3.4 whisker_0.3-2 pkgconfig_2.0.2
[43] xml2_1.2.0 lubridate_1.7.4 assertthat_0.2.0
[46] rmarkdown_1.10 httr_1.3.1 rstudioapi_0.8
[49] R6_2.3.0 nlme_3.1-137 git2r_0.23.0
[52] compiler_3.5.1
This reproducible R Markdown analysis was created with workflowr 1.1.1